ARFS - Using GPU#
You can leverage the GPU implementation of lightGBM (or other GBM flavours) but this often requires to compile or install some libraries or kit (such as CUDA)
[1]:
# from IPython.core.display import display, HTML
# display(HTML("<style>.container { width:95% !important; }</style>"))
import time
import numpy as np
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
from lightgbm import LGBMRegressor
import arfs
from arfs.feature_selection import GrootCV, Leshy
from arfs.utils import load_data
from arfs.benchmark import highlight_tick
rng = np.random.RandomState(seed=42)
# import warnings
# warnings.filterwarnings('ignore')
GrootCV on GPU#
If the data is small, using a GPU mught not be the most efficient.
[2]:
from sklearn.datasets import make_regression
from sklearn.model_selection import train_test_split
# Generate synthetic data with Poisson-distributed target variable
bias = 1
n_samples = 100_00 #1_000_000
n_features = 100
n_informative = 20
X, y, true_coef = make_regression(
n_samples=n_samples,
n_features=n_features,
n_informative=n_informative,
noise=1,
random_state=8,
bias=bias,
coef=True,
)
y = (y - y.mean()) / y.std()
y = np.exp(y) # Transform to positive values for Poisson distribution
y = np.random.poisson(y) # Add Poisson noise to the target variable
# dummy sample weight (e.g. exposure), smallest being 30 days
w = np.random.uniform(30 / 365, 1, size=len(y))
# make the count a Poisson rate (frequency)
y = y / w
X = pd.DataFrame(X)
X.columns = [f"pred_{i}" for i in range(X.shape[1])]
# Split the data into training and testing sets
X_train, X_test, y_train, y_test, w_train, w_test = train_test_split(
X, y, w, test_size=0.5, random_state=42
)
true_coef = pd.Series(true_coef)
true_coef.index = X.columns
true_coef = pd.Series({**{"intercept": bias}, **true_coef})
true_coef
genuine_predictors = true_coef[true_coef > 0.0]
print(f"The true coefficient of the linear data generating process are:\n {true_coef}")
The true coefficient of the linear data generating process are:
intercept 1.000000
pred_0 0.000000
pred_1 0.000000
pred_2 4.880441
pred_3 0.000000
...
pred_95 0.000000
pred_96 0.000000
pred_97 82.010316
pred_98 0.000000
pred_99 0.000000
Length: 101, dtype: float64
GPU
[3]:
%%time
feat_selector = GrootCV(
objective="rmse",
cutoff=1,
n_folds=3,
n_iter=3,
silent=True,
fastshap=True,
n_jobs=0,
lgbm_params={"device": "gpu", "gpu_device_id": 1},
)
feat_selector.fit(X_train, y_train, sample_weight=None)
print(f"The selected features: {feat_selector.get_feature_names_out()}")
print(f"The agnostic ranking: {feat_selector.ranking_}")
print(f"The naive ranking: {feat_selector.ranking_absolutes_}")
fig = feat_selector.plot_importance(n_feat_per_inch=5)
# highlight synthetic random variable
for name in true_coef.index:
if name in genuine_predictors.index:
fig = highlight_tick(figure=fig, str_match=name, color="green")
else:
fig = highlight_tick(figure=fig, str_match=name)
plt.show()
The selected features: ['pred_6' 'pred_11' 'pred_29' 'pred_34' 'pred_49' 'pred_55' 'pred_61'
'pred_62' 'pred_64' 'pred_68' 'pred_75' 'pred_81' 'pred_84' 'pred_87'
'pred_93' 'pred_97']
The agnostic ranking: [1 1 1 1 1 1 2 1 1 1 1 2 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 2 1 1 1 1 2 1 1
1 1 1 1 1 1 1 1 1 1 1 1 2 1 1 1 1 1 2 1 1 1 1 1 2 2 1 2 1 1 1 2 1 1 1 1 1
1 2 1 1 1 1 1 2 1 1 2 1 1 2 1 1 1 1 1 2 1 1 1 2 1 1]
The naive ranking: ['pred_87', 'pred_29', 'pred_62', 'pred_97', 'pred_84', 'pred_75', 'pred_68', 'pred_11', 'pred_49', 'pred_55', 'pred_81', 'pred_64', 'pred_6', 'pred_93', 'pred_61', 'pred_34', 'pred_57', 'pred_76', 'pred_39', 'pred_91', 'pred_14', 'pred_25', 'pred_0', 'pred_37', 'pred_50', 'pred_5', 'pred_52', 'pred_58', 'pred_70', 'pred_48', 'pred_54', 'pred_78', 'pred_12', 'pred_27', 'pred_65', 'pred_53', 'pred_46', 'pred_47', 'pred_86', 'pred_90', 'pred_60', 'pred_9', 'pred_31', 'pred_99', 'pred_56', 'pred_88', 'pred_38', 'pred_7', 'pred_10', 'pred_74', 'pred_19', 'pred_15', 'pred_21', 'pred_40', 'pred_2', 'pred_98', 'pred_16', 'pred_67', 'pred_26', 'pred_66', 'pred_3', 'pred_71', 'pred_43', 'pred_42', 'pred_63', 'pred_13', 'pred_45', 'pred_77', 'pred_24', 'pred_85', 'pred_4', 'pred_32', 'pred_72', 'pred_82', 'pred_22', 'pred_80', 'pred_73', 'pred_30', 'pred_92', 'pred_20', 'pred_94', 'pred_96', 'pred_69', 'pred_44', 'pred_79', 'pred_8', 'pred_1', 'pred_83', 'pred_36', 'pred_41', 'pred_18', 'pred_35', 'pred_59', 'pred_17', 'pred_28', 'pred_51', 'pred_89', 'pred_95', 'pred_33', 'pred_23']
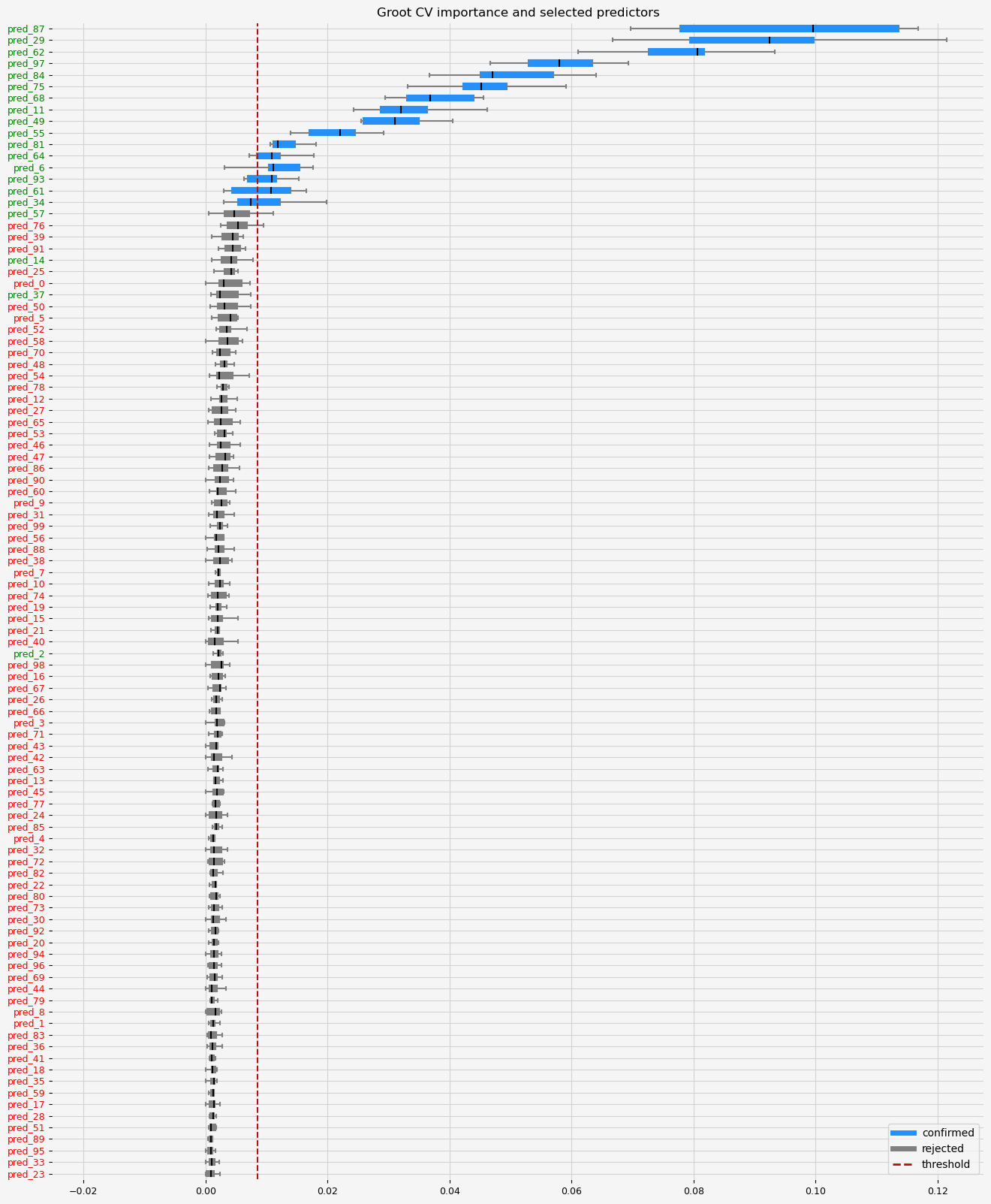
CPU times: total: 2min 38s
Wall time: 50.1 s
CPU
[4]:
%%time
feat_selector = GrootCV(
objective="rmse",
cutoff=1,
n_folds=3,
n_iter=3,
silent=True,
fastshap=True,
n_jobs=0,
lgbm_params={"device": "cpu"},
)
feat_selector.fit(X_train, y_train, sample_weight=None)
print(f"The selected features: {feat_selector.get_feature_names_out()}")
print(f"The agnostic ranking: {feat_selector.ranking_}")
print(f"The naive ranking: {feat_selector.ranking_absolutes_}")
fig = feat_selector.plot_importance(n_feat_per_inch=5)
# highlight synthetic random variable
for name in true_coef.index:
if name in genuine_predictors.index:
fig = highlight_tick(figure=fig, str_match=name, color="green")
else:
fig = highlight_tick(figure=fig, str_match=name)
plt.show()
The selected features: ['pred_6' 'pred_11' 'pred_29' 'pred_34' 'pred_49' 'pred_55' 'pred_61'
'pred_62' 'pred_64' 'pred_68' 'pred_75' 'pred_81' 'pred_84' 'pred_87'
'pred_93' 'pred_97']
The agnostic ranking: [1 1 1 1 1 1 2 1 1 1 1 2 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 2 1 1 1 1 2 1 1
1 1 1 1 1 1 1 1 1 1 1 1 2 1 1 1 1 1 2 1 1 1 1 1 2 2 1 2 1 1 1 2 1 1 1 1 1
1 2 1 1 1 1 1 2 1 1 2 1 1 2 1 1 1 1 1 2 1 1 1 2 1 1]
The naive ranking: ['pred_87', 'pred_29', 'pred_62', 'pred_97', 'pred_84', 'pred_75', 'pred_68', 'pred_11', 'pred_49', 'pred_55', 'pred_81', 'pred_6', 'pred_64', 'pred_93', 'pred_61', 'pred_34', 'pred_57', 'pred_76', 'pred_50', 'pred_91', 'pred_39', 'pred_37', 'pred_14', 'pred_0', 'pred_5', 'pred_25', 'pred_58', 'pred_65', 'pred_86', 'pred_48', 'pred_52', 'pred_53', 'pred_78', 'pred_47', 'pred_88', 'pred_70', 'pred_27', 'pred_99', 'pred_9', 'pred_7', 'pred_60', 'pred_16', 'pred_21', 'pred_15', 'pred_74', 'pred_46', 'pred_54', 'pred_82', 'pred_56', 'pred_32', 'pred_31', 'pred_22', 'pred_12', 'pred_2', 'pred_90', 'pred_40', 'pred_26', 'pred_94', 'pred_42', 'pred_66', 'pred_38', 'pred_45', 'pred_72', 'pred_10', 'pred_67', 'pred_98', 'pred_63', 'pred_19', 'pred_18', 'pred_59', 'pred_13', 'pred_30', 'pred_17', 'pred_85', 'pred_3', 'pred_92', 'pred_8', 'pred_71', 'pred_4', 'pred_69', 'pred_28', 'pred_35', 'pred_77', 'pred_80', 'pred_95', 'pred_44', 'pred_24', 'pred_43', 'pred_51', 'pred_1', 'pred_73', 'pred_89', 'pred_41', 'pred_20', 'pred_33', 'pred_96', 'pred_83', 'pred_36', 'pred_79', 'pred_23']
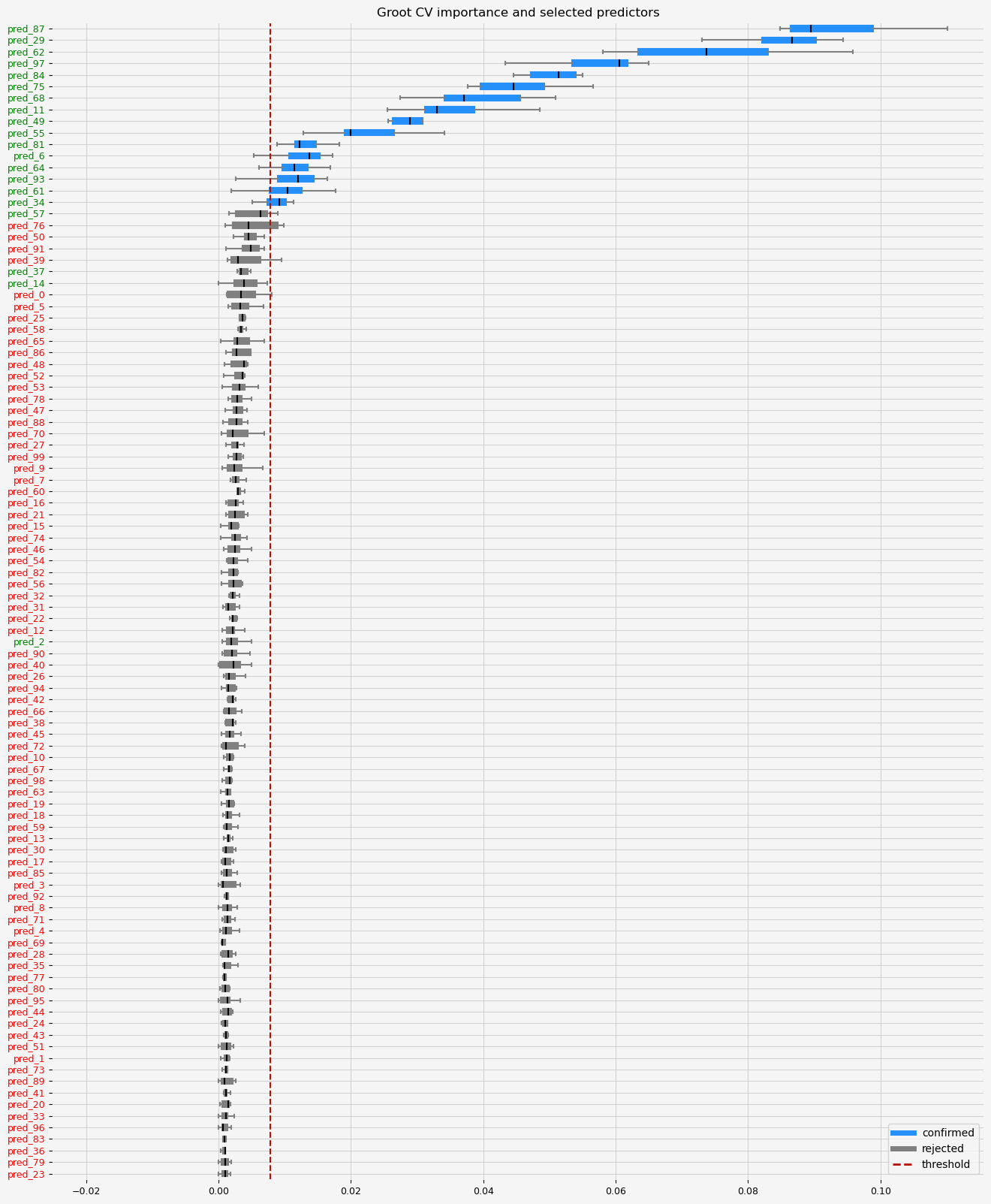
CPU times: total: 1min 18s
Wall time: 32.1 s
On a smaller data set, for illustrative purposes.
[5]:
boston = load_data(name="Boston")
X, y = boston.data, boston.target
[6]:
%%time
feat_selector = GrootCV(
objective="rmse",
cutoff=1,
n_folds=5,
n_iter=5,
silent=True,
fastshap=True,
n_jobs=0,
lgbm_params={"device": "cpu"},
)
feat_selector.fit(X, y, sample_weight=None)
print(f"The selected features: {feat_selector.get_feature_names_out()}")
print(f"The agnostic ranking: {feat_selector.ranking_}")
print(f"The naive ranking: {feat_selector.ranking_absolutes_}")
fig = feat_selector.plot_importance(n_feat_per_inch=5)
# highlight synthetic random variable
fig = highlight_tick(figure=fig, str_match="random")
fig = highlight_tick(figure=fig, str_match="genuine", color="green")
plt.show()
The selected features: ['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'TAX' 'PTRATIO' 'B' 'LSTAT' 'genuine_num']
The agnostic ranking: [2 1 1 1 2 2 2 2 1 2 2 2 2 1 1 1 1 2]
The naive ranking: ['LSTAT', 'RM', 'genuine_num', 'PTRATIO', 'DIS', 'CRIM', 'NOX', 'AGE', 'TAX', 'B', 'random_num1', 'INDUS', 'random_cat', 'random_cat_2', 'RAD', 'ZN', 'random_num2', 'CHAS']
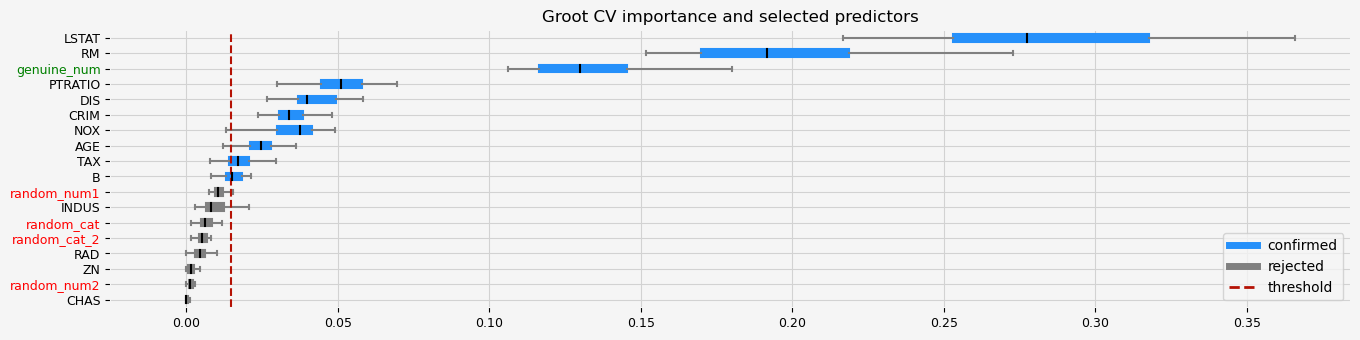
CPU times: total: 1min 2s
Wall time: 20.1 s
[7]:
%%time
feat_selector = GrootCV(
objective="rmse",
cutoff=1,
n_folds=5,
n_iter=5,
silent=True,
fastshap=True,
n_jobs=0,
lgbm_params={"device": "gpu"},
)
feat_selector.fit(X, y, sample_weight=None)
print(f"The selected features: {feat_selector.get_feature_names_out()}")
print(f"The agnostic ranking: {feat_selector.ranking_}")
print(f"The naive ranking: {feat_selector.ranking_absolutes_}")
fig = feat_selector.plot_importance(n_feat_per_inch=5)
# highlight synthetic random variable
fig = highlight_tick(figure=fig, str_match="random")
fig = highlight_tick(figure=fig, str_match="genuine", color="green")
plt.show()
The selected features: ['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'TAX' 'PTRATIO' 'LSTAT' 'genuine_num']
The agnostic ranking: [2 1 1 1 2 2 2 2 1 2 2 1 2 1 1 1 1 2]
The naive ranking: ['LSTAT', 'RM', 'genuine_num', 'PTRATIO', 'DIS', 'CRIM', 'NOX', 'AGE', 'TAX', 'B', 'random_num1', 'INDUS', 'random_cat', 'random_cat_2', 'RAD', 'ZN', 'random_num2', 'CHAS']
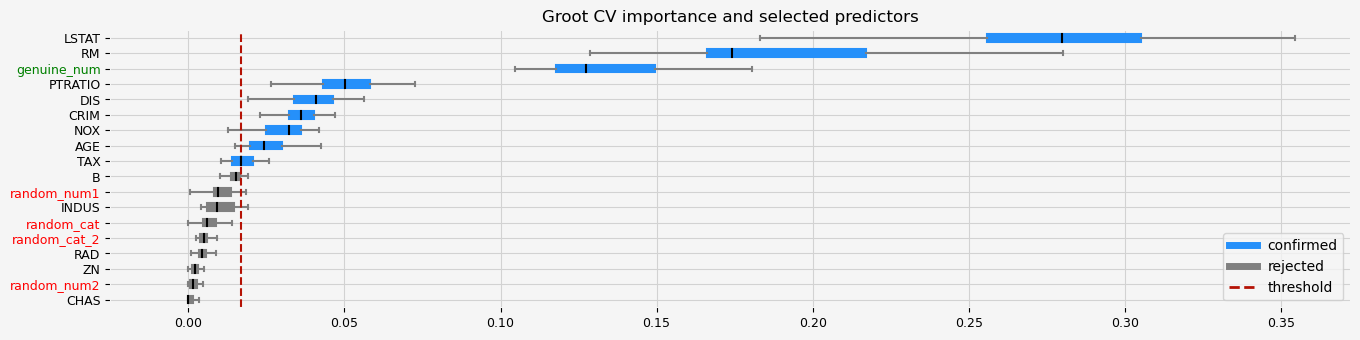
CPU times: total: 3min 55s
Wall time: 52.3 s
[8]:
%%time
feat_selector = GrootCV(
objective="rmse",
cutoff=1,
n_folds=5,
n_iter=5,
silent=True,
fastshap=True,
n_jobs=0,
lgbm_params={"device": "cuda"},
)
feat_selector.fit(X, y, sample_weight=None)
print(f"The selected features: {feat_selector.get_feature_names_out()}")
print(f"The agnostic ranking: {feat_selector.ranking_}")
print(f"The naive ranking: {feat_selector.ranking_absolutes_}")
fig = feat_selector.plot_importance(n_feat_per_inch=5)
# highlight synthetic random variable
fig = highlight_tick(figure=fig, str_match="random")
fig = highlight_tick(figure=fig, str_match="genuine", color="green")
plt.show()
---------------------------------------------------------------------------
LightGBMError Traceback (most recent call last)
File <timed exec>:11
File ~\OneDrive - Allianz\Documents\Projects\GitHub-TB\allrelevantfs\src\arfs\feature_selection\allrelevant.py:2056, in GrootCV.fit(self, X, y, sample_weight)
2053 # internal encoding (ordinal encoding)
2054 X, obj_feat, cat_idx = get_pandas_cat_codes(X)
-> 2056 self.selected_features_, self.cv_df, self.sha_cutoff = _reduce_vars_lgb_cv(
2057 X,
2058 y,
2059 objective=self.objective,
2060 cutoff=self.cutoff,
2061 n_folds=self.n_folds,
2062 n_iter=self.n_iter,
2063 silent=self.silent,
2064 weight=sample_weight,
2065 rf=self.rf,
2066 fastshap=self.fastshap,
2067 lgbm_params=self.lgbm_params,
2068 n_jobs=self.n_jobs,
2069 )
2071 self.selected_features_ = self.selected_features_.values
2072 self.support_ = np.asarray(
2073 [c in self.selected_features_ for c in self.feature_names_in_]
2074 )
File ~\OneDrive - Allianz\Documents\Projects\GitHub-TB\allrelevantfs\src\arfs\feature_selection\allrelevant.py:2270, in _reduce_vars_lgb_cv(X, y, objective, n_folds, cutoff, n_iter, silent, weight, rf, fastshap, lgbm_params, n_jobs)
2267 new_x_tr, shadow_names = _create_shadow(X_train)
2268 new_x_val, _ = _create_shadow(X_val)
-> 2270 bst, shap_matrix, bst.best_iteration = _train_lgb_model(
2271 new_x_tr,
2272 y_train,
2273 weight_tr,
2274 new_x_val,
2275 y_val,
2276 weight_val,
2277 category_cols=category_cols,
2278 early_stopping_rounds=20,
2279 fastshap=fastshap,
2280 **params,
2281 )
2283 importance = _compute_importance(
2284 new_x_tr, shap_matrix, params, objective, fastshap
2285 )
2286 df = _merge_importance_df(
2287 df=df,
2288 importance=importance,
(...)
2292 silent=silent,
2293 )
File ~\OneDrive - Allianz\Documents\Projects\GitHub-TB\allrelevantfs\src\arfs\feature_selection\allrelevant.py:2511, in _train_lgb_model(X_train, y_train, weight_train, X_val, y_val, weight_val, category_cols, early_stopping_rounds, fastshap, **params)
2506 d_valid = lgb.Dataset(
2507 X_val, label=y_val, weight=weight_val, categorical_feature=category_cols
2508 )
2509 watchlist = [d_train, d_valid]
-> 2511 bst = lgb.train(
2512 params,
2513 train_set=d_train,
2514 num_boost_round=10000,
2515 valid_sets=watchlist,
2516 categorical_feature=category_cols,
2517 callbacks=[early_stopping(early_stopping_rounds, False, False)],
2518 )
2520 if fastshap:
2521 try:
File c:\Users\xtbury\AppData\Local\mambaforge\envs\arfs\lib\site-packages\lightgbm\engine.py:271, in train(params, train_set, num_boost_round, valid_sets, valid_names, fobj, feval, init_model, feature_name, categorical_feature, early_stopping_rounds, evals_result, verbose_eval, learning_rates, keep_training_booster, callbacks)
269 # construct booster
270 try:
--> 271 booster = Booster(params=params, train_set=train_set)
272 if is_valid_contain_train:
273 booster.set_train_data_name(train_data_name)
File c:\Users\xtbury\AppData\Local\mambaforge\envs\arfs\lib\site-packages\lightgbm\basic.py:2610, in Booster.__init__(self, params, train_set, model_file, model_str, silent)
2608 params_str = param_dict_to_str(params)
2609 self.handle = ctypes.c_void_p()
-> 2610 _safe_call(_LIB.LGBM_BoosterCreate(
2611 train_set.handle,
2612 c_str(params_str),
2613 ctypes.byref(self.handle)))
2614 # save reference to data
2615 self.train_set = train_set
File c:\Users\xtbury\AppData\Local\mambaforge\envs\arfs\lib\site-packages\lightgbm\basic.py:125, in _safe_call(ret)
117 """Check the return value from C API call.
118
119 Parameters
(...)
122 The return value from C API calls.
123 """
124 if ret != 0:
--> 125 raise LightGBMError(_LIB.LGBM_GetLastError().decode('utf-8'))
LightGBMError: CUDA Tree Learner was not enabled in this build.
Please recompile with CMake option -DUSE_CUDA=1
Leshy on GPU#
[9]:
model = LGBMRegressor(random_state=42, verbose=-1, device="gpu")
[10]:
%%time
# Leshy
feat_selector = Leshy(
model, n_estimators=20, verbose=1, max_iter=10, random_state=42, importance="native"
)
feat_selector.fit(X, y, sample_weight=None)
print(f"The selected features: {feat_selector.get_feature_names_out()}")
print(f"The agnostic ranking: {feat_selector.ranking_}")
print(f"The naive ranking: {feat_selector.ranking_absolutes_}")
fig = feat_selector.plot_importance(n_feat_per_inch=5)
# highlight synthetic random variable
fig = highlight_tick(figure=fig, str_match="random")
fig = highlight_tick(figure=fig, str_match="genuine", color="green")
plt.show()
Leshy finished running using native var. imp.
Iteration: 1 / 10
Confirmed: 9
Tentative: 2
Rejected: 7
All relevant predictors selected in 00:00:02.63
The selected features: ['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'PTRATIO' 'B' 'LSTAT' 'genuine_num']
The agnostic ranking: [1 7 3 8 1 1 1 1 4 2 1 1 1 2 7 3 5 1]
The naive ranking: ['RM', 'genuine_num', 'LSTAT', 'CRIM', 'NOX', 'DIS', 'AGE', 'PTRATIO', 'B', 'TAX', 'random_num1', 'INDUS', 'random_cat', 'RAD', 'random_cat_2', 'random_num2', 'ZN', 'CHAS']
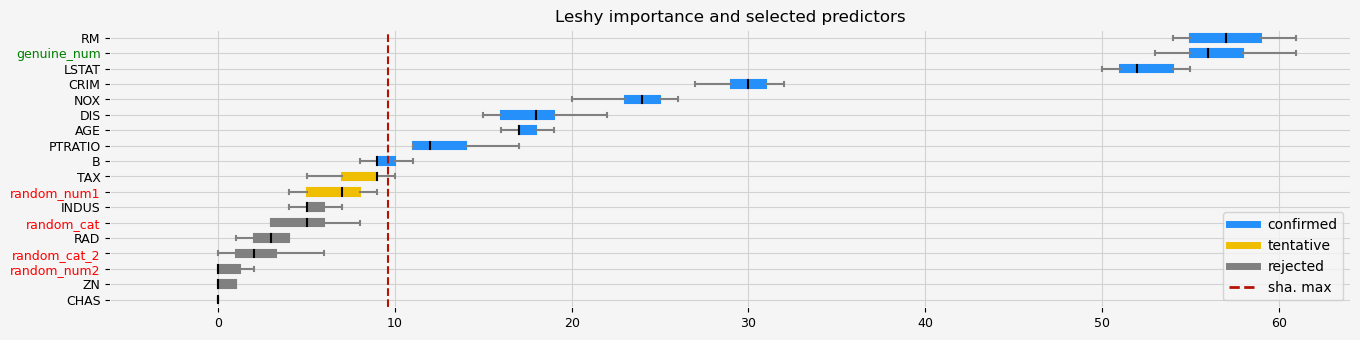
CPU times: total: 10.9 s
Wall time: 3.3 s
[11]:
model = LGBMRegressor(random_state=42, verbose=-1, device="cpu")
[12]:
%%time
# Leshy
feat_selector = Leshy(
model, n_estimators=20, verbose=1, max_iter=10, random_state=42, importance="native"
)
feat_selector.fit(X, y, sample_weight=None)
print(f"The selected features: {feat_selector.get_feature_names_out()}")
print(f"The agnostic ranking: {feat_selector.ranking_}")
print(f"The naive ranking: {feat_selector.ranking_absolutes_}")
fig = feat_selector.plot_importance(n_feat_per_inch=5)
# highlight synthetic random variable
fig = highlight_tick(figure=fig, str_match="random")
fig = highlight_tick(figure=fig, str_match="genuine", color="green")
plt.show()
Leshy finished running using native var. imp.
Iteration: 1 / 10
Confirmed: 9
Tentative: 2
Rejected: 7
All relevant predictors selected in 00:00:00.44
The selected features: ['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'PTRATIO' 'B' 'LSTAT' 'genuine_num']
The agnostic ranking: [1 7 3 8 1 1 1 1 4 2 1 1 1 2 7 3 5 1]
The naive ranking: ['RM', 'genuine_num', 'LSTAT', 'CRIM', 'NOX', 'DIS', 'AGE', 'PTRATIO', 'B', 'TAX', 'random_num1', 'INDUS', 'random_cat', 'RAD', 'random_cat_2', 'random_num2', 'ZN', 'CHAS']
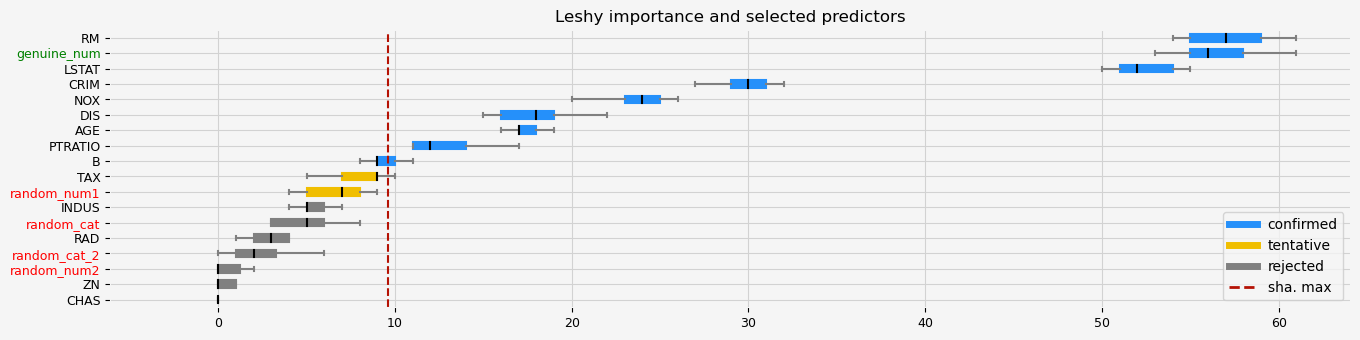
CPU times: total: 2.97 s
Wall time: 1.16 s