ARFS - regression#
ARFS can be used for classification (binary or multi-class) and for regression. You just have to specify the right loss function.
[1]:
# from IPython.core.display import display, HTML
# display(HTML("<style>.container { width:95% !important; }</style>"))
import catboost
import numpy as np
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
import gc
import shap
from boruta import BorutaPy as bp
from sklearn.datasets import fetch_openml
from sklearn.inspection import permutation_importance
from sklearn.pipeline import Pipeline
from sklearn.datasets import fetch_openml
from sklearn.inspection import permutation_importance
from sklearn.base import clone
from sklearn.ensemble import RandomForestRegressor, RandomForestClassifier
from lightgbm import LGBMRegressor, LGBMClassifier
from xgboost import XGBRegressor, XGBClassifier
from catboost import CatBoostRegressor, CatBoostClassifier
from sys import getsizeof, path
import arfs
import arfs.feature_selection as arfsfs
import arfs.feature_selection.allrelevant as arfsgroot
from arfs.feature_selection import (
MinRedundancyMaxRelevance,
GrootCV,
MissingValueThreshold,
UniqueValuesThreshold,
CollinearityThreshold,
make_fs_summary,
)
from arfs.utils import LightForestClassifier, LightForestRegressor
from arfs.benchmark import highlight_tick, compare_varimp, sklearn_pimp_bench
from arfs.utils import load_data
plt.style.use("fivethirtyeight")
rng = np.random.RandomState(seed=42)
# import warnings
# warnings.filterwarnings('ignore')
Using `tqdm.autonotebook.tqdm` in notebook mode. Use `tqdm.tqdm` instead to force console mode (e.g. in jupyter console)
[2]:
print(f"Run with ARFS {arfs.__version__}")
Run with ARFS 2.2.3
[3]:
%matplotlib inline
[4]:
gc.enable()
gc.collect()
[4]:
4
Simple Usage#
In the following examples, I’ll use a classical data set to which I added random predictors (numerical and categorical). An All Relveant FS methods should discard them. In the unit tests, you’ll find examples using artifical data with genuine (correlated and non-linear) predictors and with some random/noise columns.
Leshy (Boruta evolution)#
[5]:
boston = load_data(name="Boston")
X, y = boston.data, boston.target
[6]:
X.dtypes
[6]:
CRIM float64
ZN float64
INDUS float64
CHAS category
NOX float64
RM float64
AGE float64
DIS float64
RAD category
TAX float64
PTRATIO float64
B float64
LSTAT float64
random_num1 float64
random_num2 int32
random_cat category
random_cat_2 category
genuine_num float64
dtype: object
[7]:
X.head()
[7]:
| CRIM | ZN | INDUS | CHAS | NOX | RM | AGE | DIS | RAD | TAX | PTRATIO | B | LSTAT | random_num1 | random_num2 | random_cat | random_cat_2 | genuine_num | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 0.00632 | 18.0 | 2.31 | 0.0 | 0.538 | 6.575 | 65.2 | 4.0900 | 1.0 | 296.0 | 15.3 | 396.90 | 4.98 | 0.496714 | 0 | cat_3517 | Platist | 7.080332 |
| 1 | 0.02731 | 0.0 | 7.07 | 0.0 | 0.469 | 6.421 | 78.9 | 4.9671 | 2.0 | 242.0 | 17.8 | 396.90 | 9.14 | -0.138264 | 0 | cat_2397 | MarkZ | 5.245384 |
| 2 | 0.02729 | 0.0 | 7.07 | 0.0 | 0.469 | 7.185 | 61.1 | 4.9671 | 2.0 | 242.0 | 17.8 | 392.83 | 4.03 | 0.647689 | 0 | cat_3735 | Dracula | 6.375795 |
| 3 | 0.03237 | 0.0 | 2.18 | 0.0 | 0.458 | 6.998 | 45.8 | 6.0622 | 3.0 | 222.0 | 18.7 | 394.63 | 2.94 | 1.523030 | 0 | cat_2870 | Bejita | 6.725118 |
| 4 | 0.06905 | 0.0 | 2.18 | 0.0 | 0.458 | 7.147 | 54.2 | 6.0622 | 3.0 | 222.0 | 18.7 | 396.90 | 5.33 | -0.234153 | 4 | cat_1160 | Variance | 7.867781 |
[8]:
# Let's use lightgbm as booster, see below for using more models
model = LGBMRegressor(random_state=42, verbose=-1)
Native (impurity/Gini) feature importance, known to be biased.
[9]:
%%time
# Leshy
feat_selector = arfsgroot.Leshy(
model, n_estimators=20, verbose=1, max_iter=10, random_state=42, importance="native"
)
feat_selector.fit(X, y, sample_weight=None)
print(f"The selected features: {feat_selector.get_feature_names_out()}")
print(f"The agnostic ranking: {feat_selector.ranking_}")
print(f"The naive ranking: {feat_selector.ranking_absolutes_}")
fig = feat_selector.plot_importance(n_feat_per_inch=5)
# highlight synthetic random variable
fig = highlight_tick(figure=fig, str_match="random")
fig = highlight_tick(figure=fig, str_match="genuine", color="green")
plt.show()
Leshy finished running using native var. imp.
Iteration: 1 / 10
Confirmed: 9
Tentative: 2
Rejected: 7
All relevant predictors selected in 00:00:00.83
The selected features: ['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'PTRATIO' 'B' 'LSTAT' 'genuine_num']
The agnostic ranking: [1 7 3 8 1 1 1 1 4 2 1 1 1 2 7 3 5 1]
The naive ranking: ['RM', 'genuine_num', 'LSTAT', 'CRIM', 'NOX', 'DIS', 'AGE', 'PTRATIO', 'B', 'TAX', 'random_num1', 'INDUS', 'random_cat', 'RAD', 'random_cat_2', 'random_num2', 'ZN', 'CHAS']
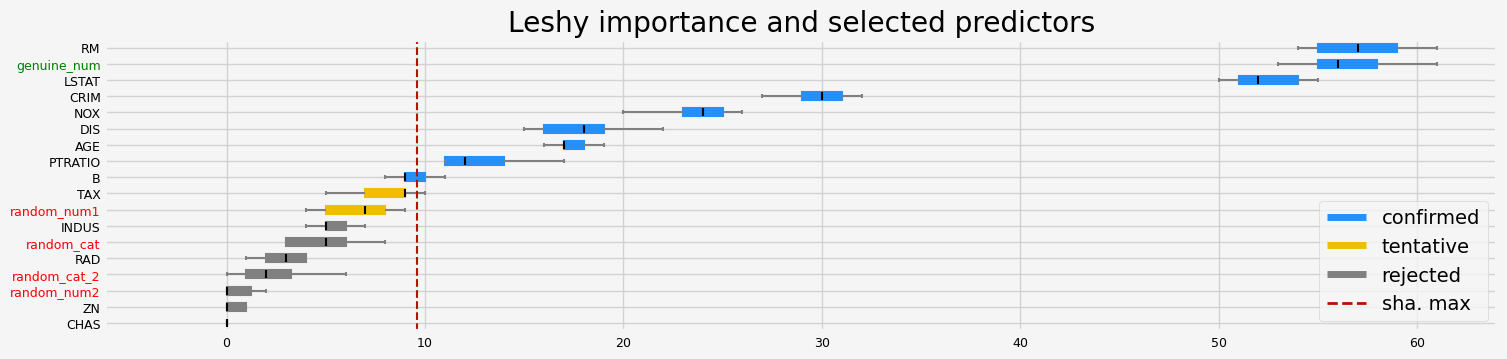
CPU times: user 2.98 s, sys: 241 ms, total: 3.22 s
Wall time: 1.5 s
SHAP importance
[10]:
%%time
model = clone(model)
# Leshy
feat_selector = arfsgroot.Leshy(
model, n_estimators=20, verbose=1, max_iter=10, random_state=42, importance="shap"
)
feat_selector.fit(X, y, sample_weight=None)
print(f"The selected features: {feat_selector.get_feature_names_out()}")
print(f"The agnostic ranking: {feat_selector.ranking_}")
print(f"The naive ranking: {feat_selector.ranking_absolutes_}")
fig = feat_selector.plot_importance(n_feat_per_inch=5)
# highlight synthetic random variable
fig = highlight_tick(figure=fig, str_match="random")
fig = highlight_tick(figure=fig, str_match="genuine", color="green")
plt.show()
Leshy finished running using shap var. imp.
Iteration: 1 / 10
Confirmed: 9
Tentative: 1
Rejected: 8
All relevant predictors selected in 00:00:00.54
The selected features: ['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'PTRATIO' 'LSTAT' 'random_num1'
'genuine_num']
The agnostic ranking: [1 9 4 9 1 1 1 1 6 2 1 3 1 1 9 5 6 1]
The naive ranking: ['LSTAT', 'RM', 'genuine_num', 'CRIM', 'PTRATIO', 'DIS', 'AGE', 'random_num1', 'NOX', 'TAX', 'B', 'INDUS', 'random_cat', 'random_cat_2', 'RAD', 'random_num2', 'ZN', 'CHAS']
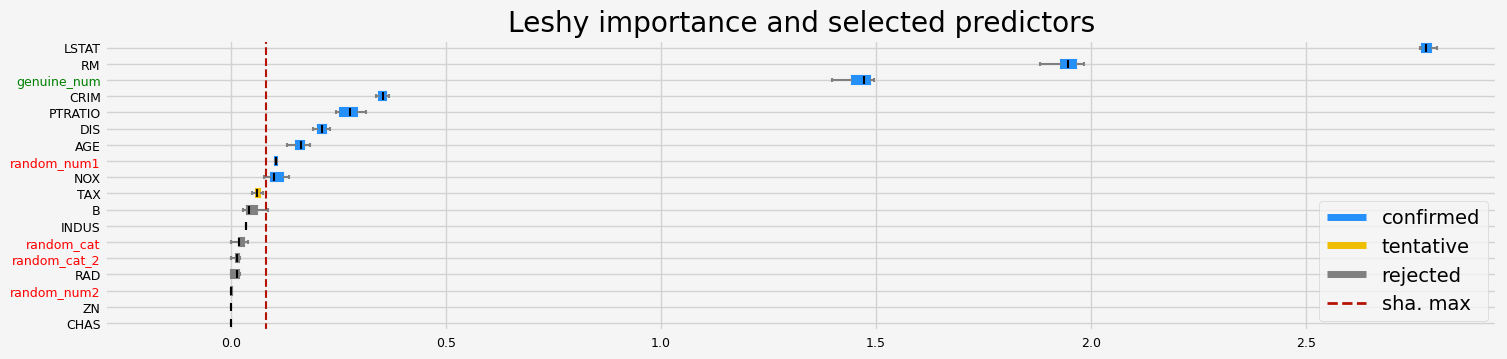
CPU times: user 2.39 s, sys: 239 ms, total: 2.63 s
Wall time: 1.24 s
SHAP importance - fasttreeshap implementation
[11]:
%%time
model = clone(model)
# Leshy
feat_selector = arfsgroot.Leshy(
model,
n_estimators=20,
verbose=1,
max_iter=10,
random_state=42,
importance="fastshap",
)
feat_selector.fit(X, y, sample_weight=None)
print(f"The selected features: {feat_selector.get_feature_names_out()}")
print(f"The agnostic ranking: {feat_selector.ranking_}")
print(f"The naive ranking: {feat_selector.ranking_absolutes_}")
fig = feat_selector.plot_importance(n_feat_per_inch=5)
# highlight synthetic random variable
fig = highlight_tick(figure=fig, str_match="random")
fig = highlight_tick(figure=fig, str_match="genuine", color="green")
plt.show()
Leshy finished running using native var. imp.
Iteration: 1 / 10
Confirmed: 9
Tentative: 1
Rejected: 8
All relevant predictors selected in 00:00:00.55
The selected features: ['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'PTRATIO' 'LSTAT' 'random_num1'
'genuine_num']
The agnostic ranking: [1 9 4 9 1 1 1 1 6 2 1 3 1 1 9 5 6 1]
The naive ranking: ['LSTAT', 'RM', 'genuine_num', 'CRIM', 'PTRATIO', 'DIS', 'AGE', 'random_num1', 'NOX', 'TAX', 'B', 'INDUS', 'random_cat', 'random_cat_2', 'RAD', 'random_num2', 'ZN', 'CHAS']
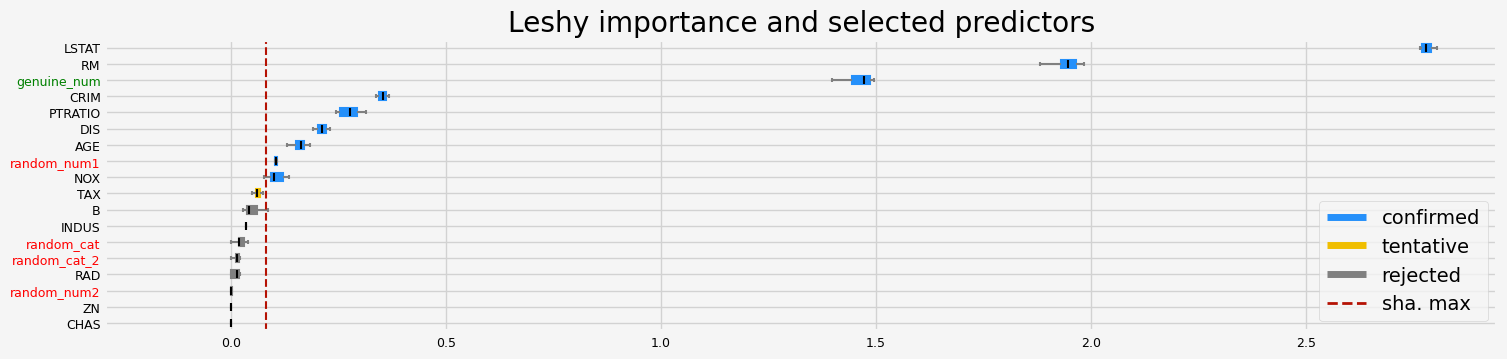
CPU times: user 2.56 s, sys: 242 ms, total: 2.8 s
Wall time: 1.26 s
with permutation importance
[12]:
%%time
model = clone(model)
# Leshy
feat_selector = arfsgroot.Leshy(
model, n_estimators=20, verbose=1, max_iter=10, random_state=42, importance="pimp"
)
feat_selector.fit(X, y, sample_weight=None)
print(f"The selected features: {feat_selector.get_feature_names_out()}")
print(f"The agnostic ranking: {feat_selector.ranking_}")
print(f"The naive ranking: {feat_selector.ranking_absolutes_}")
fig = feat_selector.plot_importance(n_feat_per_inch=5)
# highlight synthetic random variable
fig = highlight_tick(figure=fig, str_match="random")
fig = highlight_tick(figure=fig, str_match="genuine", color="green")
plt.show()
Leshy finished running using pimp var. imp.
Iteration: 1 / 10
Confirmed: 8
Tentative: 1
Rejected: 9
All relevant predictors selected in 00:00:05.30
The selected features: ['CRIM' 'NOX' 'RM' 'DIS' 'TAX' 'PTRATIO' 'LSTAT' 'genuine_num']
The agnostic ranking: [ 1 7 3 7 1 1 11 1 4 1 1 2 1 4 9 6 10 1]
The naive ranking: ['LSTAT', 'RM', 'genuine_num', 'CRIM', 'DIS', 'PTRATIO', 'NOX', 'TAX', 'B', 'RAD', 'INDUS', 'random_num1', 'CHAS', 'ZN', 'random_num2', 'AGE', 'random_cat', 'random_cat_2']
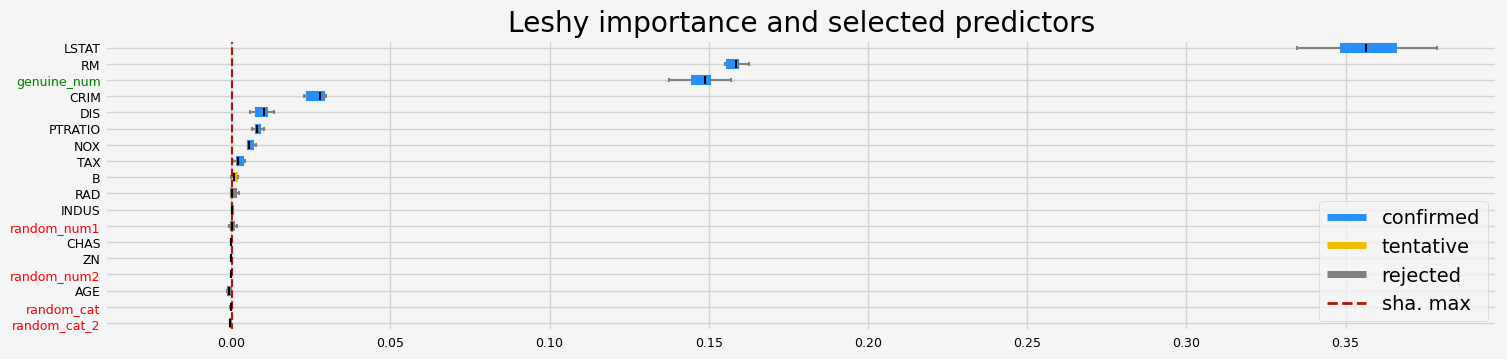
CPU times: user 2.94 s, sys: 479 ms, total: 3.42 s
Wall time: 5.93 s
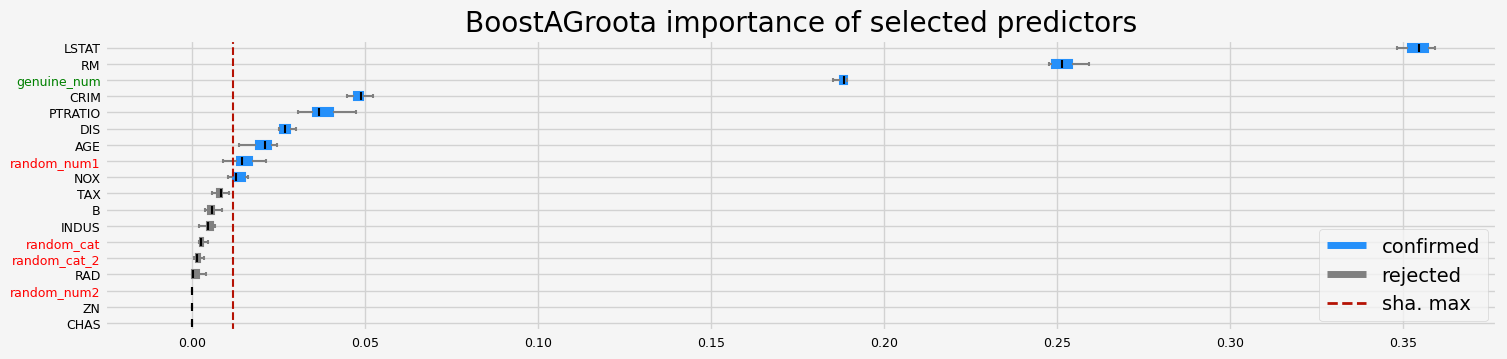
BoostAGroota#
with SHAP importance
[13]:
%%time
# be sure to use the same but non-fitted estimator
model = clone(model)
# BoostAGroota
feat_selector = arfsgroot.BoostAGroota(
estimator=model, cutoff=1, iters=10, max_rounds=10, delta=0.1, importance="shap"
)
feat_selector.fit(X, y, sample_weight=None)
print(f"The selected features: {feat_selector.get_feature_names_out()}")
print(f"The agnostic ranking: {feat_selector.ranking_}")
print(f"The naive ranking: {feat_selector.ranking_absolutes_}")
fig = feat_selector.plot_importance(n_feat_per_inch=5)
# highlight synthetic random variable
fig = highlight_tick(figure=fig, str_match="random")
fig = highlight_tick(figure=fig, str_match="genuine", color="green")
plt.show()
The selected features: ['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'PTRATIO' 'LSTAT' 'random_num1'
'genuine_num']
The agnostic ranking: [2 1 1 1 2 2 2 2 1 1 2 1 2 2 1 1 1 2]
The naive ranking: ['LSTAT', 'RM', 'genuine_num', 'CRIM', 'PTRATIO', 'DIS', 'AGE', 'random_num1', 'NOX', 'TAX', 'B', 'INDUS', 'random_cat', 'random_cat_2', 'RAD', 'random_num2', 'ZN', 'CHAS']
CPU times: user 2.23 s, sys: 322 ms, total: 2.55 s
Wall time: 1.61 s
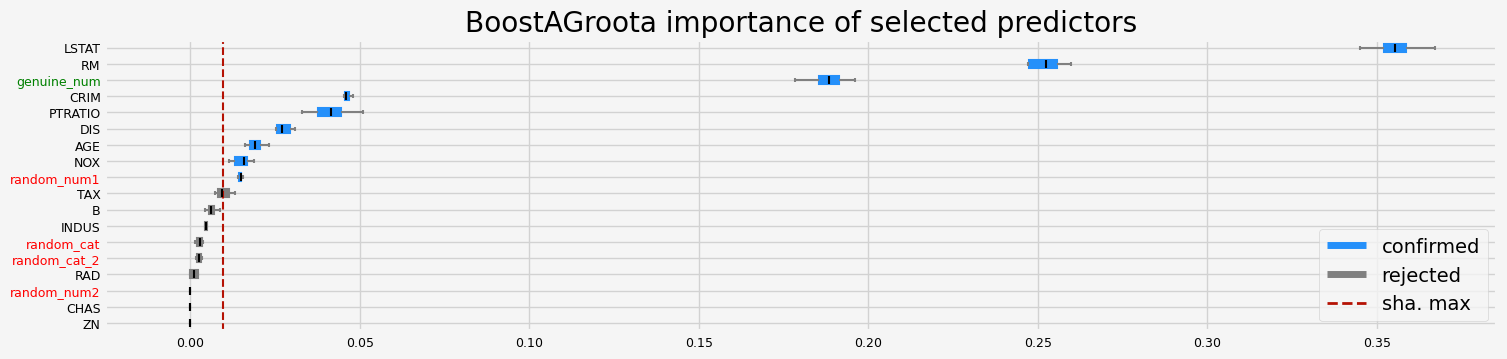
with SHAP importance - fasttreeshap implementation
[14]:
%%time
# be sure to use the same but non-fitted estimator
model = clone(model)
# BoostAGroota
feat_selector = arfsgroot.BoostAGroota(
estimator=model, cutoff=1, iters=10, max_rounds=10, delta=0.1, importance="fastshap"
)
feat_selector.fit(X, y, sample_weight=None)
print(f"The selected features: {feat_selector.get_feature_names_out()}")
print(f"The agnostic ranking: {feat_selector.ranking_}")
print(f"The naive ranking: {feat_selector.ranking_absolutes_}")
fig = feat_selector.plot_importance(n_feat_per_inch=5)
# highlight synthetic random variable
fig = highlight_tick(figure=fig, str_match="random")
fig = highlight_tick(figure=fig, str_match="genuine", color="green")
plt.show()
The selected features: ['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'PTRATIO' 'LSTAT' 'random_num1'
'genuine_num']
The agnostic ranking: [2 1 1 1 2 2 2 2 1 1 2 1 2 2 1 1 1 2]
The naive ranking: ['LSTAT', 'RM', 'genuine_num', 'CRIM', 'PTRATIO', 'DIS', 'AGE', 'NOX', 'random_num1', 'TAX', 'B', 'INDUS', 'random_cat', 'random_cat_2', 'RAD', 'random_num2', 'CHAS', 'ZN']
CPU times: user 2.68 s, sys: 272 ms, total: 2.95 s
Wall time: 1.72 s
[15]:
feat_selector.get_params()
[15]:
{'cutoff': 1,
'delta': 0.1,
'estimator__boosting_type': 'gbdt',
'estimator__class_weight': None,
'estimator__colsample_bytree': 1.0,
'estimator__importance_type': 'split',
'estimator__learning_rate': 0.1,
'estimator__max_depth': -1,
'estimator__min_child_samples': 20,
'estimator__min_child_weight': 0.001,
'estimator__min_split_gain': 0.0,
'estimator__n_estimators': 20,
'estimator__n_jobs': -1,
'estimator__num_leaves': 31,
'estimator__objective': None,
'estimator__random_state': 2600,
'estimator__reg_alpha': 0.0,
'estimator__reg_lambda': 0.0,
'estimator__silent': 'warn',
'estimator__subsample': 1.0,
'estimator__subsample_for_bin': 200000,
'estimator__subsample_freq': 0,
'estimator__verbose': -1,
'estimator': LGBMRegressor(n_estimators=20, random_state=2600, verbose=-1),
'importance': 'fastshap',
'iters': 10,
'max_rounds': 10,
'silent': True}
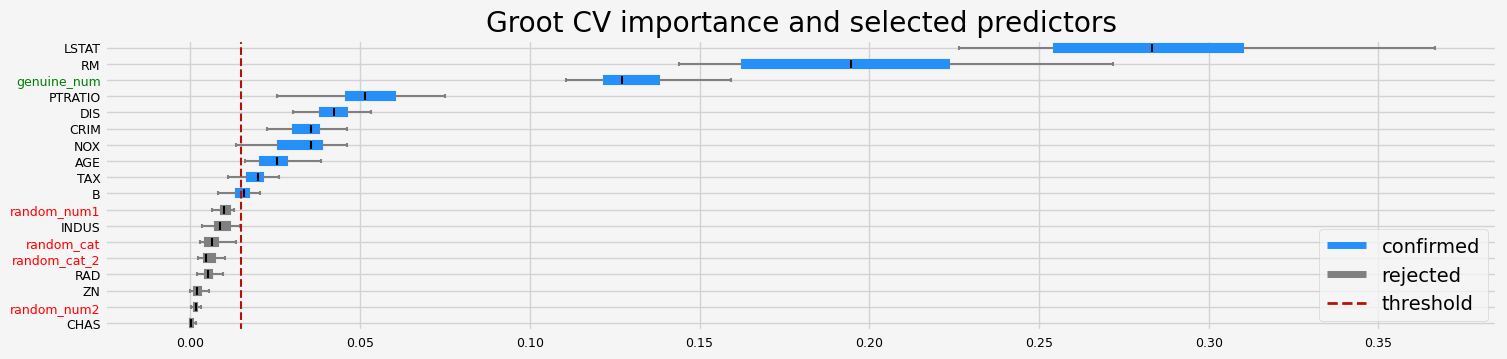
GrootCV#
[16]:
%%time
# GrootCV
feat_selector = arfsgroot.GrootCV(
objective="rmse",
cutoff=1,
n_folds=5,
n_iter=5,
silent=True,
fastshap=False,
n_jobs=0,
)
feat_selector.fit(X, y, sample_weight=None)
print(f"The selected features: {feat_selector.get_feature_names_out()}")
print(f"The agnostic ranking: {feat_selector.ranking_}")
print(f"The naive ranking: {feat_selector.ranking_absolutes_}")
fig = feat_selector.plot_importance(n_feat_per_inch=5)
# highlight synthetic random variable
fig = highlight_tick(figure=fig, str_match="random")
fig = highlight_tick(figure=fig, str_match="genuine", color="green")
plt.show()
The selected features: ['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'TAX' 'PTRATIO' 'B' 'LSTAT' 'genuine_num']
The agnostic ranking: [2 1 1 1 2 2 2 2 1 2 2 2 2 1 1 1 1 2]
The naive ranking: ['LSTAT', 'RM', 'genuine_num', 'PTRATIO', 'DIS', 'CRIM', 'NOX', 'AGE', 'TAX', 'B', 'random_num1', 'INDUS', 'random_cat', 'random_cat_2', 'RAD', 'ZN', 'random_num2', 'CHAS']
CPU times: user 24.7 s, sys: 1.78 s, total: 26.5 s
Wall time: 10.9 s
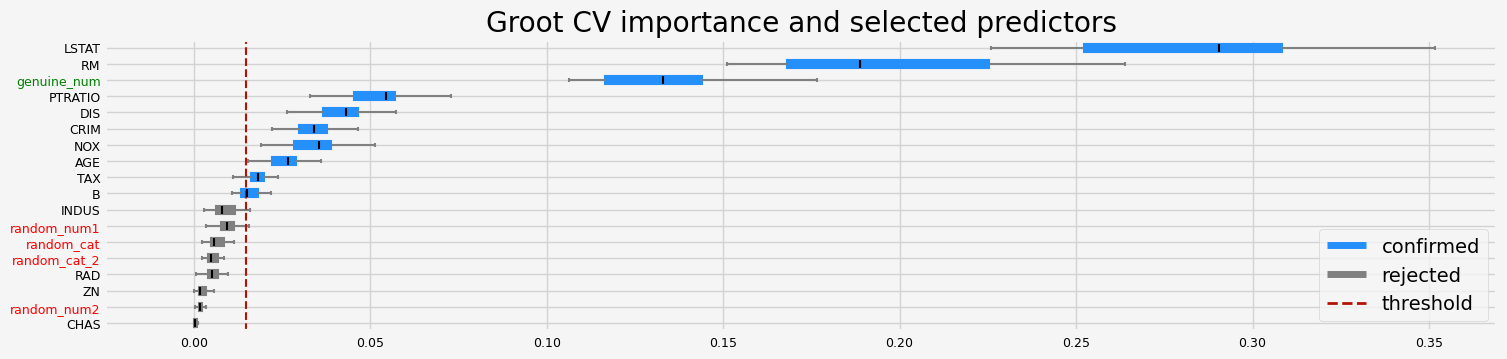
enabling fasttreeshap
[17]:
%%time
# GrootCV
feat_selector = arfsgroot.GrootCV(
objective="rmse",
cutoff=1,
n_folds=5,
n_iter=5,
silent=True,
fastshap=True,
n_jobs=0,
)
feat_selector.fit(X, y, sample_weight=None)
print(f"The selected features: {feat_selector.get_feature_names_out()}")
print(f"The agnostic ranking: {feat_selector.ranking_}")
print(f"The naive ranking: {feat_selector.ranking_absolutes_}")
fig = feat_selector.plot_importance(n_feat_per_inch=5)
# highlight synthetic random variable
fig = highlight_tick(figure=fig, str_match="random")
fig = highlight_tick(figure=fig, str_match="genuine", color="green")
plt.show()
The selected features: ['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'TAX' 'PTRATIO' 'B' 'LSTAT' 'genuine_num']
The agnostic ranking: [2 1 1 1 2 2 2 2 1 2 2 2 2 1 1 1 1 2]
The naive ranking: ['LSTAT', 'RM', 'genuine_num', 'PTRATIO', 'DIS', 'CRIM', 'NOX', 'AGE', 'TAX', 'B', 'INDUS', 'random_num1', 'random_cat', 'random_cat_2', 'RAD', 'ZN', 'random_num2', 'CHAS']
CPU times: user 24.3 s, sys: 2.12 s, total: 26.4 s
Wall time: 12.7 s
ARFS in sklearn pipelines#
all the selectors (basic, arfs and MRmr) are sklearn compatible and follows the same architecture. Namely, they use the sklearn relevant base classes and therefore have the same methods.
[18]:
feat_selector = arfsgroot.GrootCV(
objective="rmse", cutoff=1, n_folds=5, n_iter=5, silent=True
)
arfs_fs_pipeline = Pipeline(
[
("missing", MissingValueThreshold(threshold=0.05)),
("unique", UniqueValuesThreshold(threshold=1)),
("collinearity", CollinearityThreshold(threshold=0.85)),
("arfs", feat_selector),
]
)
X_trans = arfs_fs_pipeline.fit(X=X, y=y).transform(X=X)
you can access the attributes of a step as you would in any sklearn pipeline
[19]:
arfs_fs_pipeline.named_steps["collinearity"].get_feature_names_out()
[19]:
array(['CRIM', 'ZN', 'INDUS', 'CHAS', 'RM', 'AGE', 'DIS', 'RAD', 'TAX',
'PTRATIO', 'B', 'LSTAT', 'random_num1', 'random_num2',
'random_cat', 'random_cat_2', 'genuine_num'], dtype=object)
[20]:
fig = arfs_fs_pipeline.named_steps["arfs"].plot_importance()
# highlight synthetic random variable
fig = highlight_tick(figure=fig, str_match="random")
fig = highlight_tick(figure=fig, str_match="genuine", color="green")
plt.show()
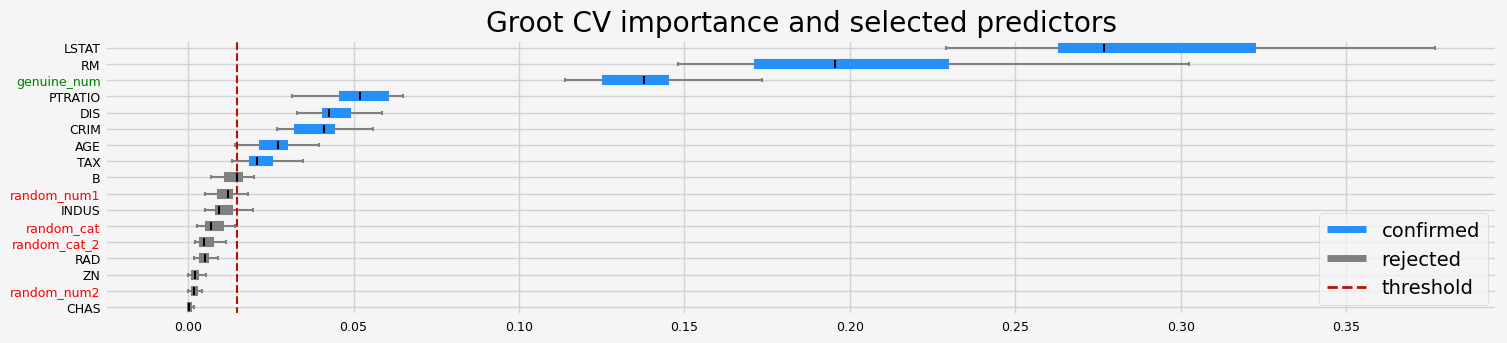
[21]:
make_fs_summary(arfs_fs_pipeline)
[21]:
| predictor | missing | unique | collinearity | arfs | |
|---|---|---|---|---|---|
| 0 | CRIM | 1 | 1 | 1 | 1 |
| 1 | ZN | 1 | 1 | 1 | 0 |
| 2 | INDUS | 1 | 1 | 1 | 0 |
| 3 | CHAS | 1 | 1 | 1 | 0 |
| 4 | NOX | 1 | 1 | 0 | nan |
| 5 | RM | 1 | 1 | 1 | 1 |
| 6 | AGE | 1 | 1 | 1 | 1 |
| 7 | DIS | 1 | 1 | 1 | 1 |
| 8 | RAD | 1 | 1 | 1 | 0 |
| 9 | TAX | 1 | 1 | 1 | 1 |
| 10 | PTRATIO | 1 | 1 | 1 | 1 |
| 11 | B | 1 | 1 | 1 | 0 |
| 12 | LSTAT | 1 | 1 | 1 | 1 |
| 13 | random_num1 | 1 | 1 | 1 | 0 |
| 14 | random_num2 | 1 | 1 | 1 | 0 |
| 15 | random_cat | 1 | 1 | 1 | 0 |
| 16 | random_cat_2 | 1 | 1 | 1 | 0 |
| 17 | genuine_num | 1 | 1 | 1 | 1 |
Testing and comparing Leshy, GrootCV and BoostAGroota#
In the following examples, I’ll use different models which are scikit-learn compatible and then one can compare the different ARFS methods with different models and the different feature importance.
[22]:
%%time
model = clone(model)
# Benchmark with scikit-learn permutation importance
print("=" * 20 + " Benchmarking using sklearn permutation importance " + "=" * 20)
fig = sklearn_pimp_bench(model, X, y, task="regression", sample_weight=None)
==================== Benchmarking using sklearn permutation importance ====================
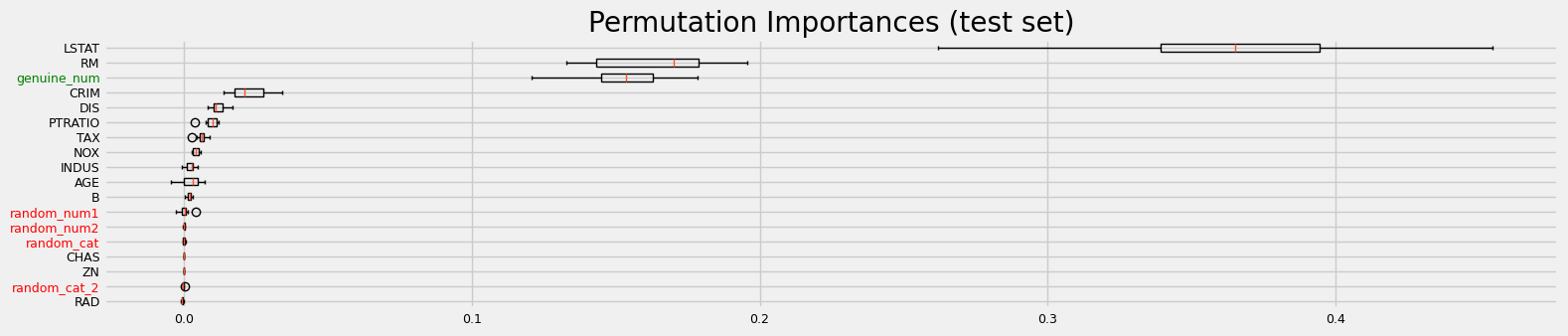
CPU times: user 862 ms, sys: 359 ms, total: 1.22 s
Wall time: 2.88 s
Testing Leshy#
Leshy seems to struggle with catboost, for regression and this particular data set whereas the other ARFS methods seem OK. To be investigated.
[23]:
models = [
RandomForestRegressor(n_jobs=4, oob_score=True),
CatBoostRegressor(random_state=42, verbose=0),
LGBMRegressor(random_state=42, verbose=-1),
LightForestRegressor(n_feat=X.shape[1]),
XGBRegressor(random_state=42, verbosity=0),
]
feat_selector = arfsgroot.Leshy(
model, n_estimators=100, verbose=1, max_iter=10, random_state=42
)
if __name__ == "__main__":
# regression
boston = load_data(name="Boston")
X, y = boston.data, boston.target
# running the ARFS methods using different models
compare_varimp(feat_selector, models, X, y, sample_weight=None)
==================== Leshy - testing: RandomForestRegressor for var.imp: shap ====================
Leshy finished running using shap var. imp.
Iteration: 1 / 10
Confirmed: 11
Tentative: 1
Rejected: 6
All relevant predictors selected in 00:00:17.14
['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'TAX' 'PTRATIO' 'B' 'LSTAT' 'random_num1'
'genuine_num']
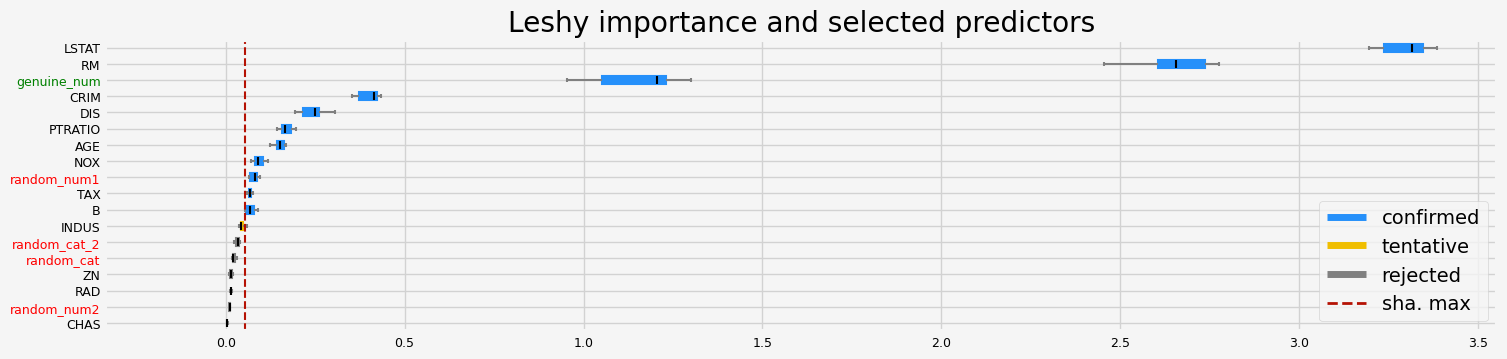
==================== Leshy - testing: RandomForestRegressor for var.imp: fastshap ====================
Leshy finished running using native var. imp.
Iteration: 1 / 10
Confirmed: 10
Tentative: 2
Rejected: 6
All relevant predictors selected in 00:00:09.47
['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'TAX' 'PTRATIO' 'LSTAT' 'random_num1'
'genuine_num']
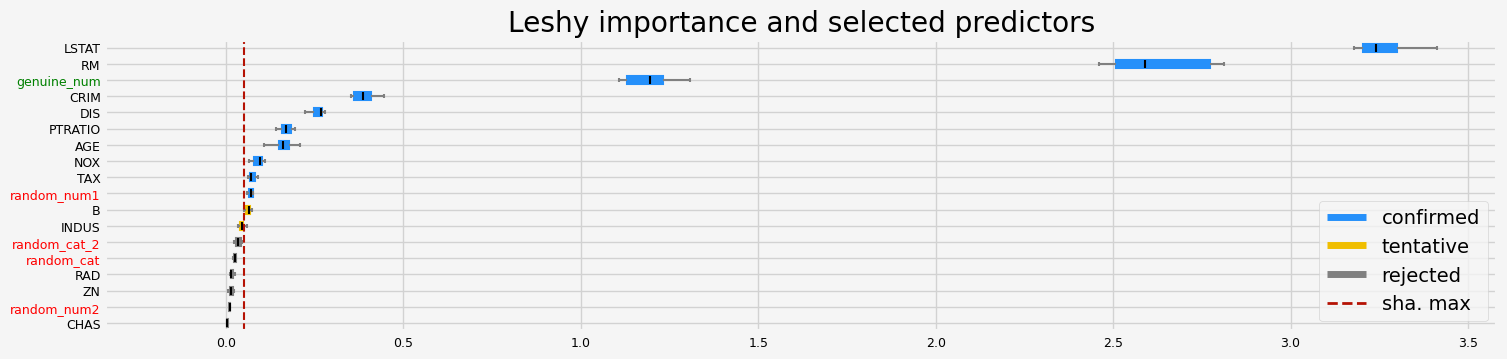
==================== Leshy - testing: RandomForestRegressor for var.imp: pimp ====================
Leshy finished running using pimp var. imp.
Iteration: 1 / 10
Confirmed: 9
Tentative: 2
Rejected: 7
All relevant predictors selected in 00:00:31.21
['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'TAX' 'PTRATIO' 'LSTAT' 'genuine_num']
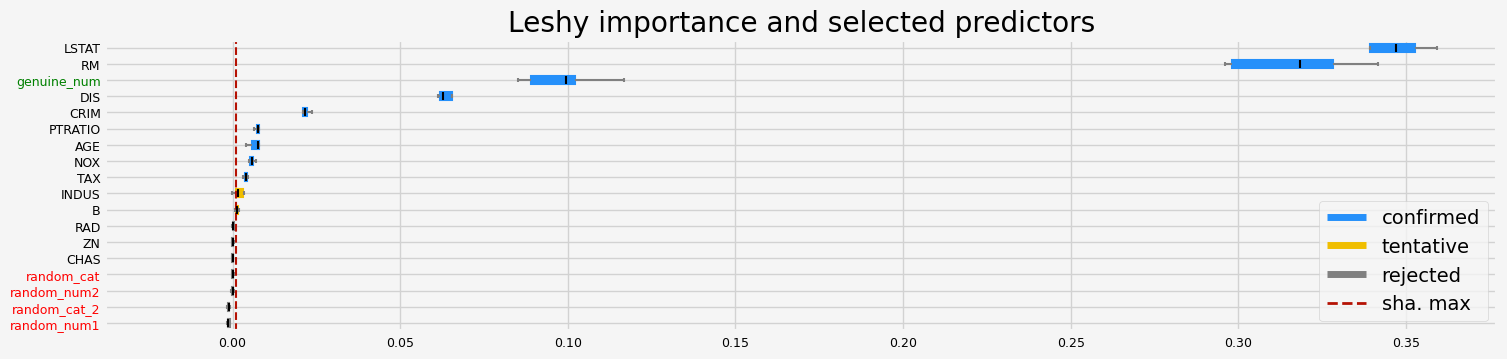
==================== Leshy - testing: RandomForestRegressor for var.imp: native ====================
Leshy finished running using native var. imp.
Iteration: 1 / 10
Confirmed: 10
Tentative: 1
Rejected: 7
All relevant predictors selected in 00:00:06.60
['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'TAX' 'PTRATIO' 'B' 'LSTAT' 'genuine_num']
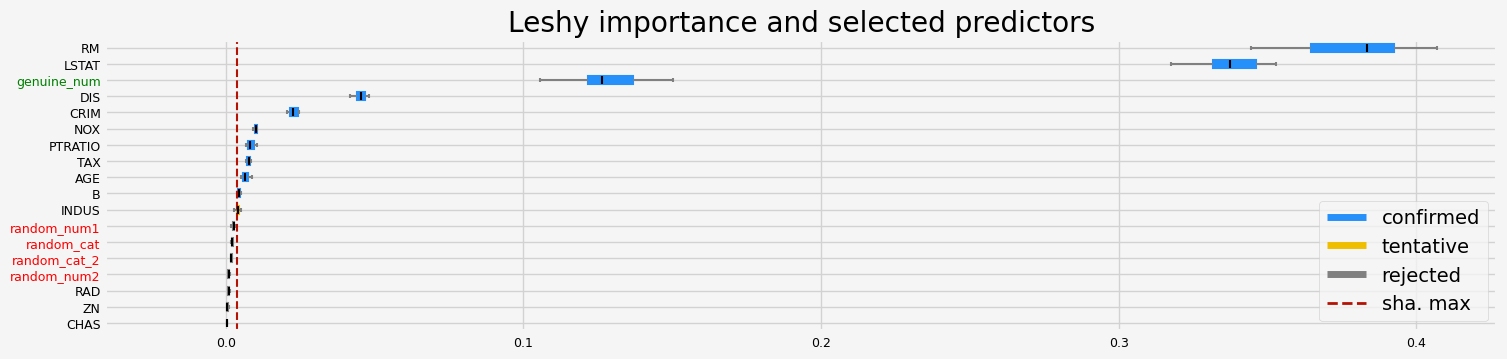
==================== Leshy - testing: CatBoostRegressor for var.imp: shap ====================
Leshy finished running using shap var. imp.
Iteration: 1 / 10
Confirmed: 10
Tentative: 3
Rejected: 5
All relevant predictors selected in 00:00:05.75
['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'TAX' 'PTRATIO' 'B' 'LSTAT' 'genuine_num']
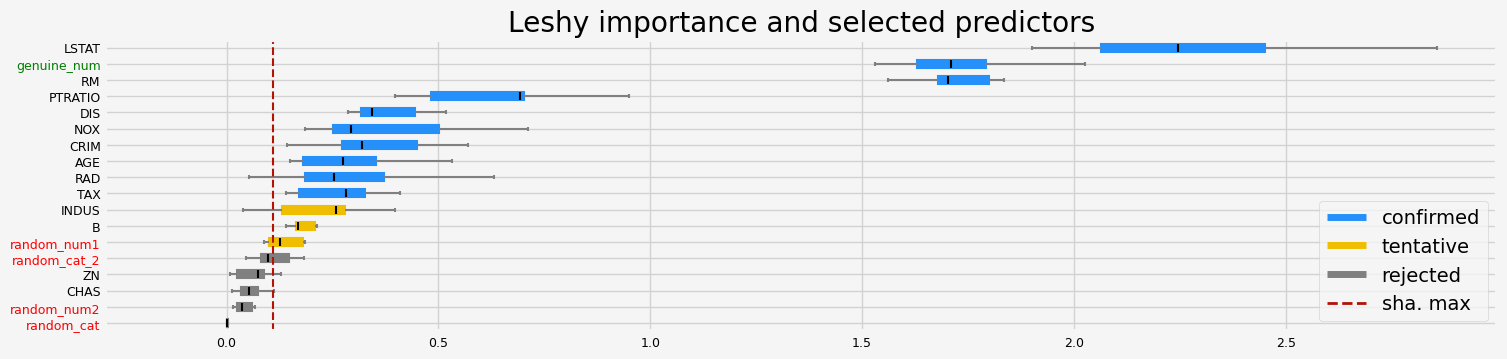
==================== Leshy - testing: CatBoostRegressor for var.imp: fastshap ====================
Leshy finished running using native var. imp.
Iteration: 1 / 10
Confirmed: 8
Tentative: 4
Rejected: 6
All relevant predictors selected in 00:00:05.44
['CRIM' 'RM' 'AGE' 'DIS' 'PTRATIO' 'B' 'LSTAT' 'genuine_num']
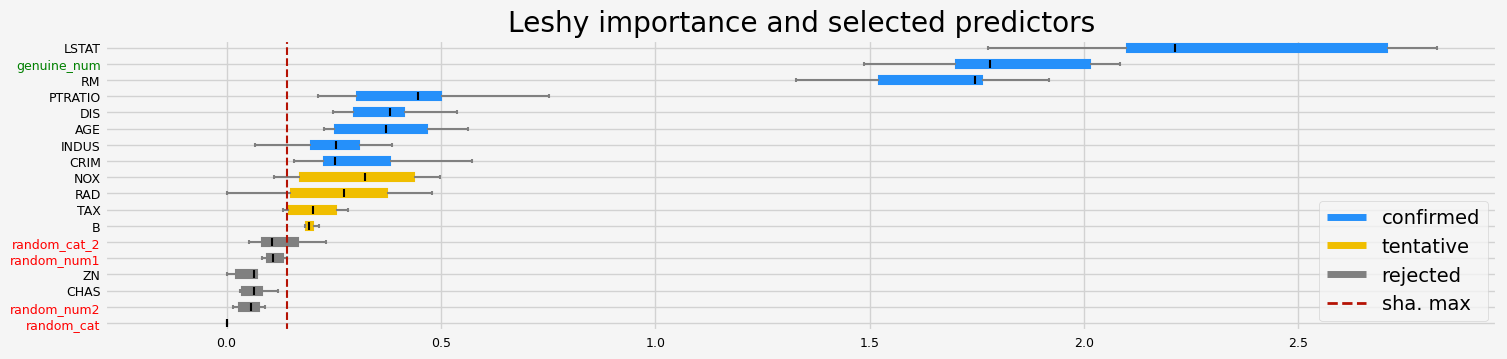
==================== Leshy - testing: CatBoostRegressor for var.imp: pimp ====================
Leshy finished running using pimp var. imp.
Iteration: 1 / 10
Confirmed: 7
Tentative: 5
Rejected: 6
All relevant predictors selected in 00:00:11.62
['RM' 'AGE' 'DIS' 'TAX' 'PTRATIO' 'LSTAT' 'genuine_num']
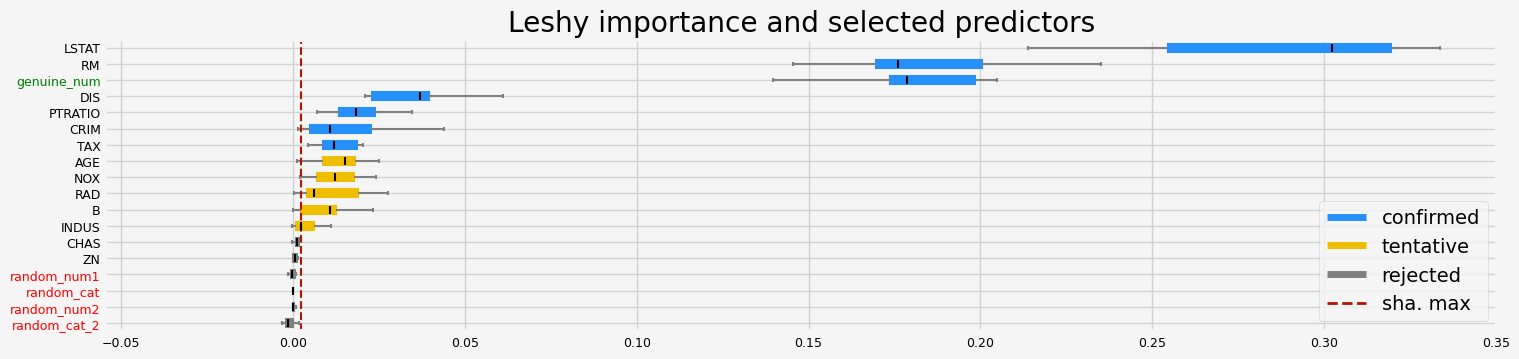
==================== Leshy - testing: CatBoostRegressor for var.imp: native ====================
Leshy finished running using native var. imp.
Iteration: 1 / 10
Confirmed: 7
Tentative: 4
Rejected: 7
All relevant predictors selected in 00:00:04.90
['CRIM' 'NOX' 'RM' 'DIS' 'PTRATIO' 'LSTAT' 'genuine_num']
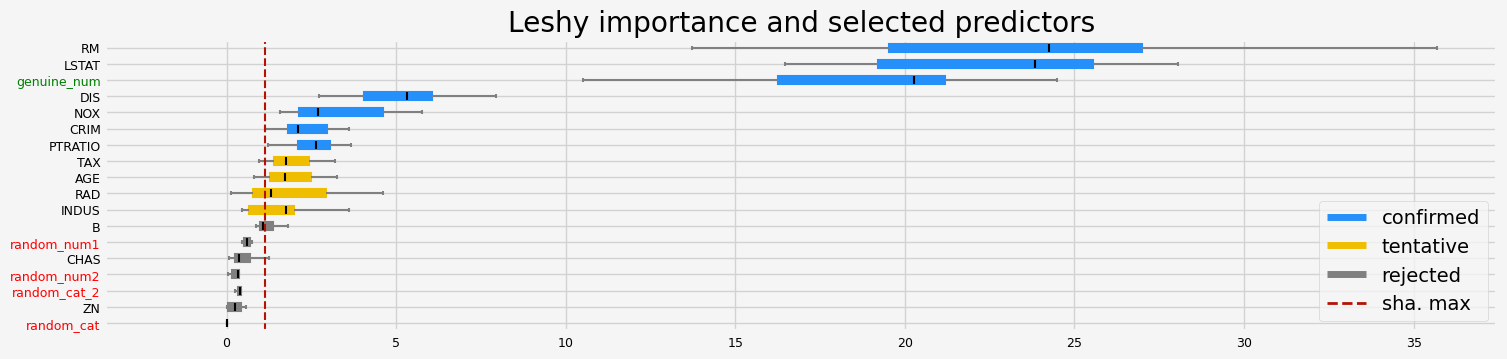
==================== Leshy - testing: LGBMRegressor for var.imp: shap ====================
Leshy finished running using shap var. imp.
Iteration: 1 / 10
Confirmed: 8
Tentative: 2
Rejected: 8
All relevant predictors selected in 00:00:02.29
['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'PTRATIO' 'LSTAT' 'genuine_num']
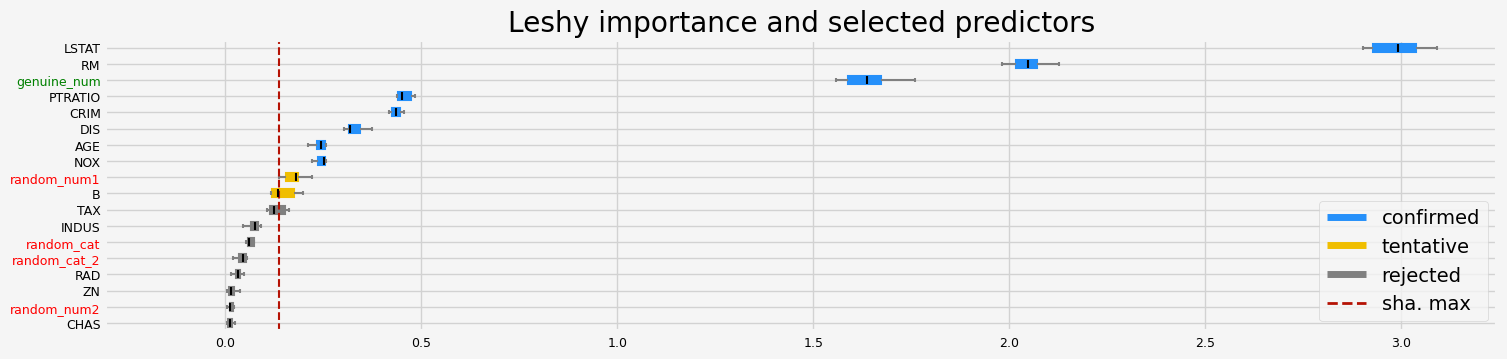
==================== Leshy - testing: LGBMRegressor for var.imp: fastshap ====================
Leshy finished running using native var. imp.
Iteration: 1 / 10
Confirmed: 8
Tentative: 0
Rejected: 10
All relevant predictors selected in 00:00:03.66
['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'PTRATIO' 'LSTAT' 'genuine_num']
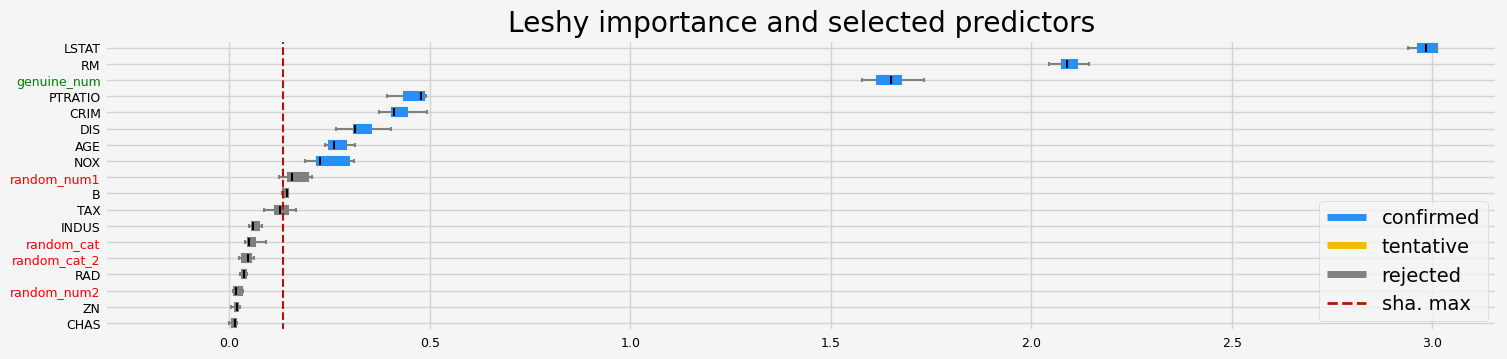
==================== Leshy - testing: LGBMRegressor for var.imp: pimp ====================
Leshy finished running using pimp var. imp.
Iteration: 1 / 10
Confirmed: 8
Tentative: 3
Rejected: 7
All relevant predictors selected in 00:00:07.78
['CRIM' 'NOX' 'RM' 'DIS' 'TAX' 'PTRATIO' 'LSTAT' 'genuine_num']
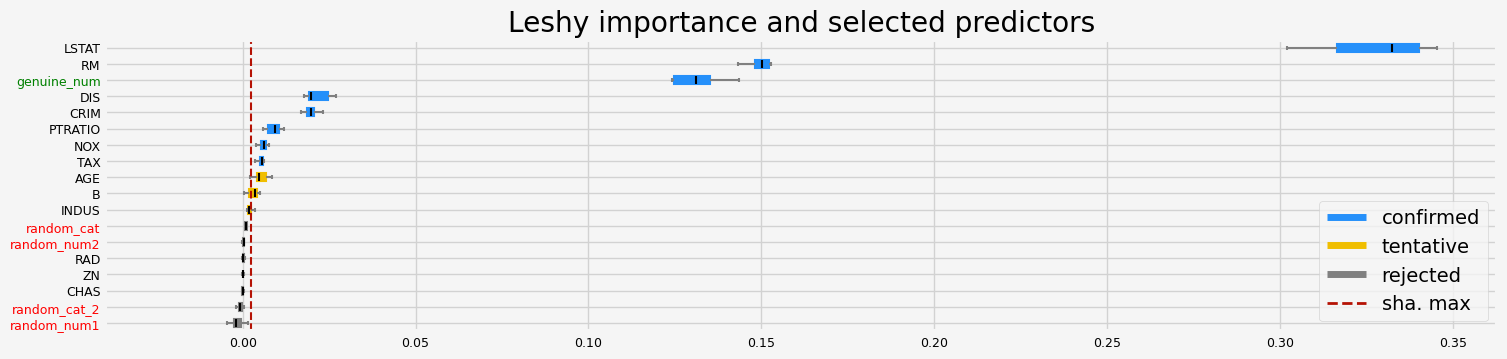
==================== Leshy - testing: LGBMRegressor for var.imp: native ====================
Leshy finished running using native var. imp.
Iteration: 1 / 10
Confirmed: 5
Tentative: 3
Rejected: 10
All relevant predictors selected in 00:00:02.70
['CRIM' 'RM' 'DIS' 'LSTAT' 'genuine_num']
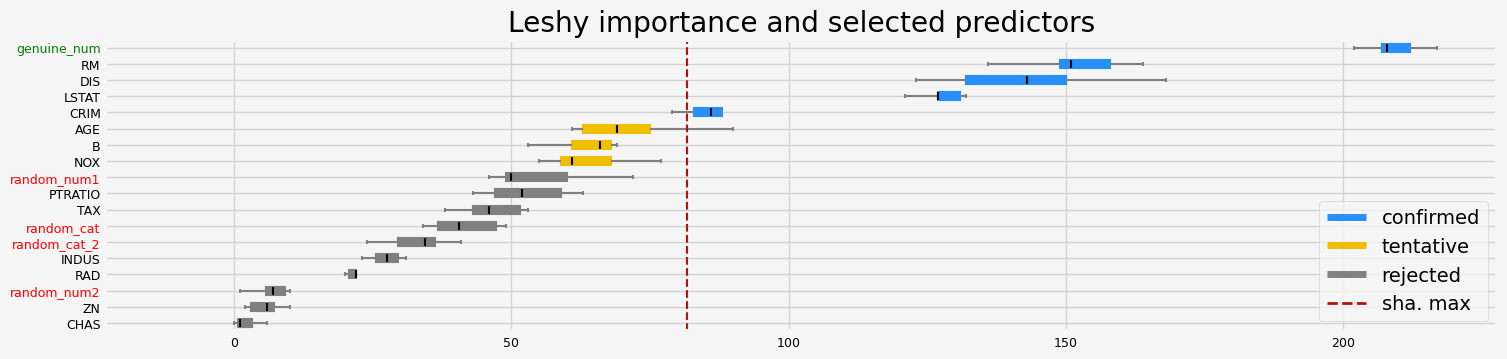
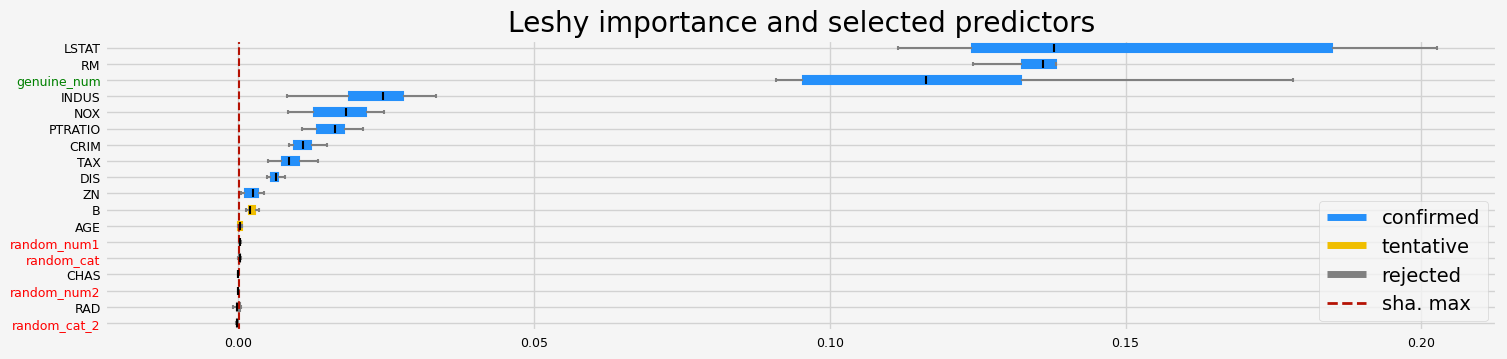
==================== Leshy - testing: LGBMRegressor for var.imp: shap ====================
Leshy finished running using shap var. imp.
Iteration: 1 / 10
Confirmed: 11
Tentative: 2
Rejected: 5
All relevant predictors selected in 00:00:01.40
['CRIM' 'INDUS' 'NOX' 'RM' 'AGE' 'DIS' 'TAX' 'PTRATIO' 'B' 'LSTAT'
'genuine_num']
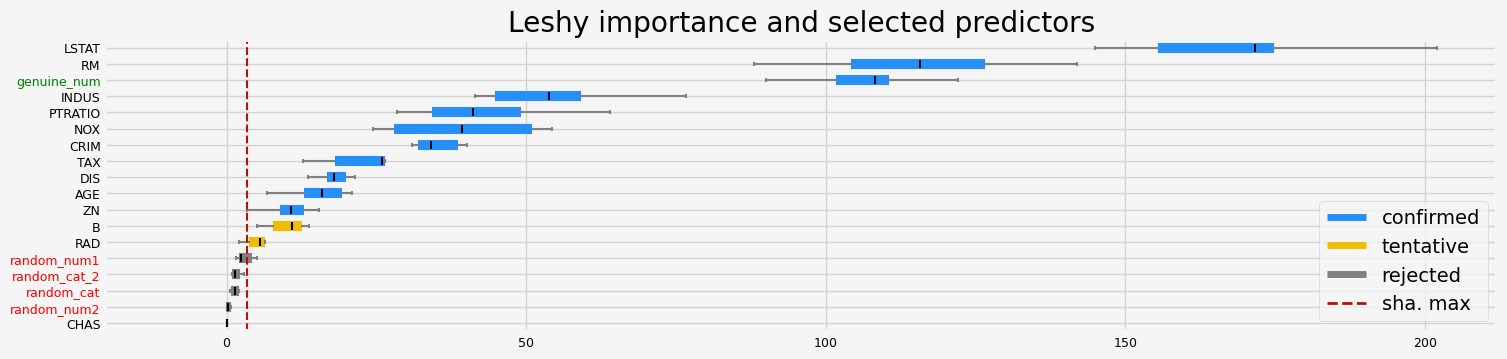
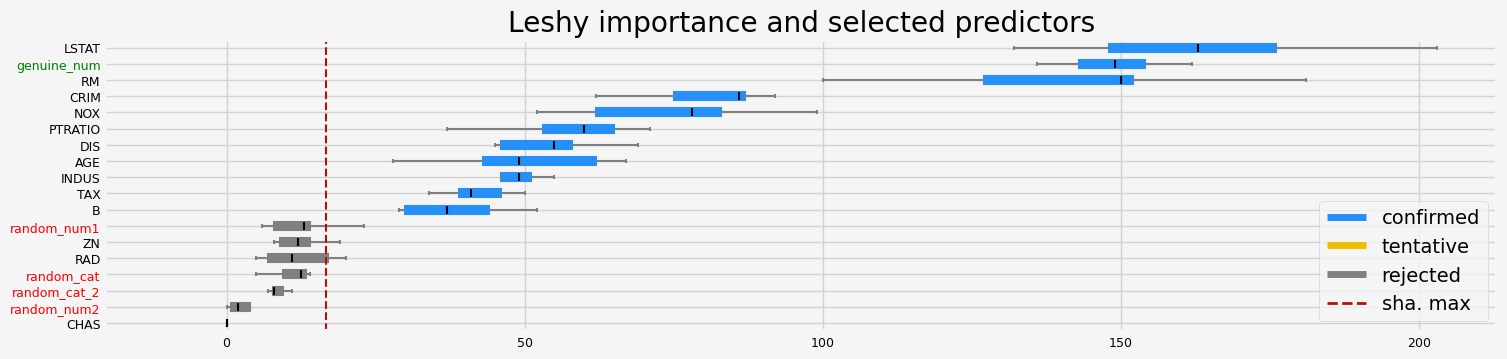
==================== Leshy - testing: LGBMRegressor for var.imp: fastshap ====================
Leshy finished running using native var. imp.
Iteration: 1 / 10
Confirmed: 12
Tentative: 1
Rejected: 5
All relevant predictors selected in 00:00:01.87
['CRIM' 'ZN' 'INDUS' 'NOX' 'RM' 'AGE' 'DIS' 'TAX' 'PTRATIO' 'B' 'LSTAT'
'genuine_num']
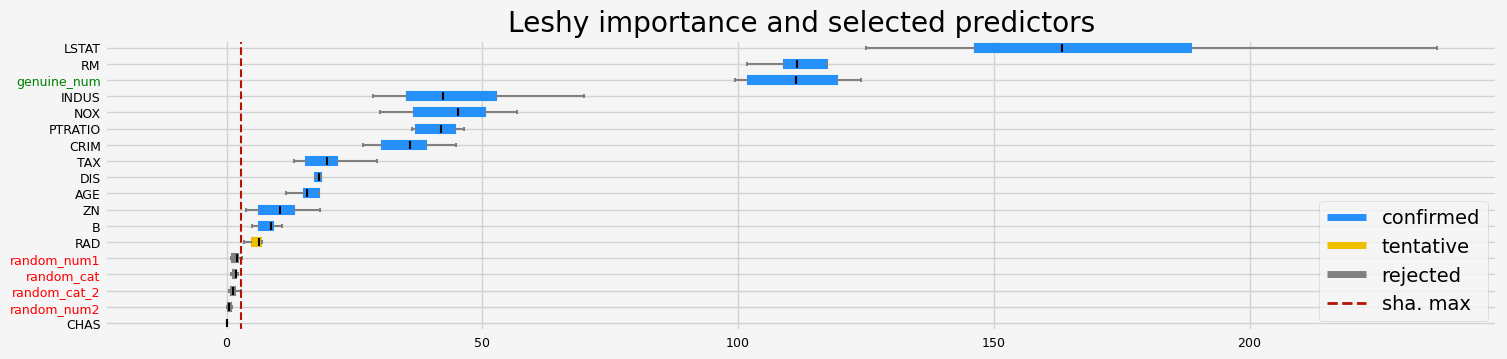
==================== Leshy - testing: LGBMRegressor for var.imp: pimp ====================
Leshy finished running using pimp var. imp.
Iteration: 1 / 10
Confirmed: 10
Tentative: 3
Rejected: 5
All relevant predictors selected in 00:00:05.26
['CRIM' 'ZN' 'INDUS' 'NOX' 'RM' 'DIS' 'TAX' 'PTRATIO' 'LSTAT'
'genuine_num']
==================== Leshy - testing: LGBMRegressor for var.imp: native ====================
Leshy finished running using native var. imp.
Iteration: 1 / 10
Confirmed: 11
Tentative: 0
Rejected: 7
All relevant predictors selected in 00:00:01.65
['CRIM' 'INDUS' 'NOX' 'RM' 'AGE' 'DIS' 'TAX' 'PTRATIO' 'B' 'LSTAT'
'genuine_num']
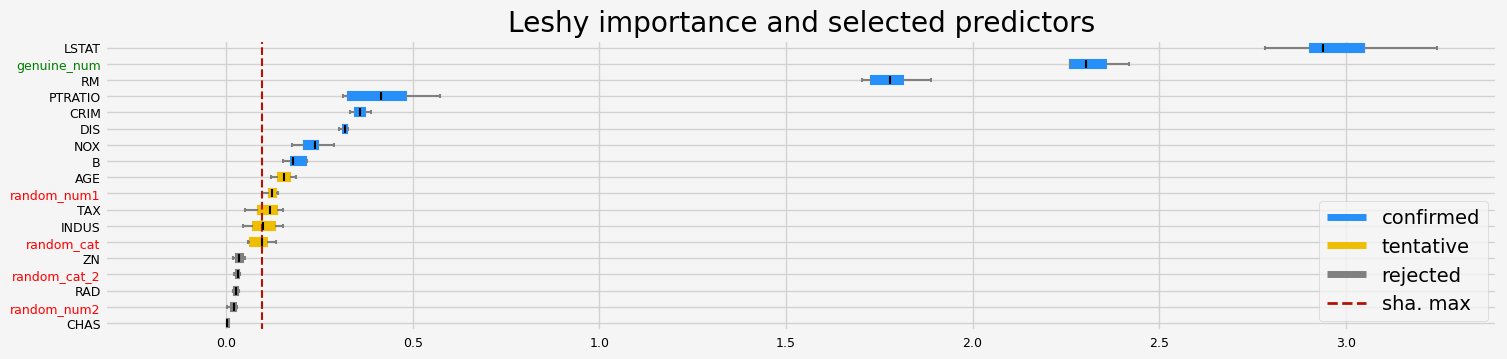
==================== Leshy - testing: XGBRegressor for var.imp: shap ====================
Leshy finished running using shap var. imp.
Iteration: 1 / 10
Confirmed: 8
Tentative: 5
Rejected: 5
All relevant predictors selected in 00:00:05.20
['CRIM' 'NOX' 'RM' 'DIS' 'PTRATIO' 'B' 'LSTAT' 'genuine_num']
==================== Leshy - testing: XGBRegressor for var.imp: fastshap ====================
---------------------------------------------------------------------------
Exception Traceback (most recent call last)
Cell In[23], line 18
16 X, y = boston.data, boston.target
17 # running the ARFS methods using different models
---> 18 compare_varimp(feat_selector, models, X, y, sample_weight=None)
File ~/Documents/arfs/src/arfs/benchmark.py:142, in compare_varimp(feat_selector, models, X, y, sample_weight)
140 feat_selector.estimator = mod_clone
141 # fit the feature selector
--> 142 feat_selector.fit(X=X, y=y, sample_weight=sample_weight)
143 # print the results
144 print(feat_selector.selected_features_)
File ~/Documents/arfs/src/arfs/feature_selection/allrelevant.py:330, in Leshy.fit(self, X, y, sample_weight)
327 raise TypeError("X is not a dataframe")
329 self.imp_real_hist = np.empty((0, X.shape[1]), float)
--> 330 self._fit(X, y, sample_weight=sample_weight)
331 self.selected_features_ = self.feature_names_in_[self.support_]
332 self.not_selected_features_ = self.feature_names_in_[~self.support_]
File ~/Documents/arfs/src/arfs/feature_selection/allrelevant.py:469, in Leshy._fit(self, X_raw, y, sample_weight)
466 if self.n_estimators != "auto":
467 self.estimator.set_params(n_estimators=self.n_estimators)
--> 469 dec_reg, sha_max_history, imp_history, imp_sha_max = self.select_features(
470 X=X, y=y, sample_weight=sample_weight
471 )
472 confirmed, tentative = _get_confirmed_and_tentative(dec_reg)
473 tentative = _select_tentative(tentative, imp_history, sha_max_history)
File ~/Documents/arfs/src/arfs/feature_selection/allrelevant.py:940, in Leshy.select_features(self, X, y, sample_weight)
932 self._update_tree_num(dec_reg)
933 self._update_estimator()
934 (
935 dec_reg,
936 sha_max_history,
937 imp_history,
938 hit_reg,
939 imp_sha_max,
--> 940 ) = self._run_iteration(
941 X,
942 y,
943 sample_weight,
944 dec_reg,
945 sha_max_history,
946 imp_history,
947 hit_reg,
948 _iter,
949 )
950 _iter += 1
951 pbar.update(1)
File ~/Documents/arfs/src/arfs/feature_selection/allrelevant.py:891, in Leshy._run_iteration(self, X, y, sample_weight, dec_reg, sha_max_history, imp_history, hit_reg, _iter)
842 def _run_iteration(
843 self, X, y, sample_weight, dec_reg, sha_max_history, imp_history, hit_reg, _iter
844 ):
845 """
846 Run an iteration of the Gradient Boosting algorithm.
847
(...)
889 The maximum shadow importance value for this iteration.
890 """
--> 891 cur_imp = self._add_shadows_get_imps(X, y, sample_weight, dec_reg)
892 imp_sha_max = np.percentile(cur_imp[1], self.perc)
893 sha_max_history.append(imp_sha_max)
File ~/Documents/arfs/src/arfs/feature_selection/allrelevant.py:558, in Leshy._add_shadows_get_imps(self, X, y, sample_weight, dec_reg)
554 imp = _get_shap_imp(
555 self.estimator, pd.concat([x_cur, x_sha], axis=1), y, sample_weight
556 )
557 elif self.importance == "fastshap":
--> 558 imp = _get_shap_imp_fast(
559 self.estimator, pd.concat([x_cur, x_sha], axis=1), y, sample_weight
560 )
561 elif self.importance == "pimp":
562 imp = _get_perm_imp(
563 self.estimator, pd.concat([x_cur, x_sha], axis=1), y, sample_weight
564 )
File ~/Documents/arfs/src/arfs/feature_selection/allrelevant.py:1302, in _get_shap_imp_fast(estimator, X, y, sample_weight, cat_feature)
1293 model, X_tt, y_tt, w_tt = _split_fit_estimator(
1294 estimator, X, y, sample_weight=sample_weight, cat_feature=cat_feature
1295 )
1296 explainer = FastTreeExplainer(
1297 model,
1298 algorithm="auto",
1299 shortcut=False,
1300 feature_perturbation="tree_path_dependent",
1301 )
-> 1302 shap_matrix = explainer.shap_values(X_tt)
1303 # multiclass returns a list
1304 # for binary and for some models, shap is still returning a list
1305 if is_classifier(estimator):
File ~/mambaforge-pypy3/envs/arfs/lib/python3.10/site-packages/fasttreeshap/explainers/_tree.py:481, in Tree.shap_values(self, X, y, tree_limit, approximate, check_additivity, from_call)
479 out = self._get_shap_output(phi, flat_output)
480 if check_additivity and self.model.model_output == "raw":
--> 481 self.assert_additivity(out, self.model.predict(X))
483 return out
File ~/mambaforge-pypy3/envs/arfs/lib/python3.10/site-packages/fasttreeshap/explainers/_tree.py:641, in Tree.assert_additivity(self, phi, model_output)
639 check_sum(self.expected_value[i] + phi[i].sum(-1), model_output[:,i])
640 else:
--> 641 check_sum(self.expected_value + phi.sum(-1), model_output)
File ~/mambaforge-pypy3/envs/arfs/lib/python3.10/site-packages/fasttreeshap/explainers/_tree.py:635, in Tree.assert_additivity.<locals>.check_sum(sum_val, model_output)
631 err_msg += " Consider retrying with the feature_perturbation='interventional' option."
632 err_msg += " This check failed because for one of the samples the sum of the SHAP values" \
633 " was %f, while the model output was %f. If this difference is acceptable" \
634 " you can set check_additivity=False to disable this check." % (sum_val[ind], model_output[ind])
--> 635 raise Exception(err_msg)
Exception: Additivity check failed in TreeExplainer! Please ensure the data matrix you passed to the explainer is the same shape that the model was trained on. If your data shape is correct then please report this on GitHub. Consider retrying with the feature_perturbation='interventional' option. This check failed because for one of the samples the sum of the SHAP values was 22.882743, while the model output was 45.290653. If this difference is acceptable you can set check_additivity=False to disable this check.
[24]:
from sklearn.datasets import make_regression
from xgboost import XGBRegressor
from lightgbm import LGBMRegressor
from fasttreeshap import TreeExplainer as FastTreeExplainer
X, y = make_regression(
n_samples=1000, n_features=10, n_informative=8, noise=1, random_state=8
)
model = XGBRegressor() # LGBMRegressor()
model.fit(X, y)
explainer = FastTreeExplainer(
model, algorithm="auto", shortcut=False, feature_perturbation="tree_path_dependent"
)
shap_matrix = explainer.shap_values(X)
---------------------------------------------------------------------------
Exception Traceback (most recent call last)
Cell In[24], line 14
10 model.fit(X, y)
11 explainer = FastTreeExplainer(
12 model, algorithm="auto", shortcut=False, feature_perturbation="tree_path_dependent"
13 )
---> 14 shap_matrix = explainer.shap_values(X)
File ~/mambaforge-pypy3/envs/arfs/lib/python3.10/site-packages/fasttreeshap/explainers/_tree.py:481, in Tree.shap_values(self, X, y, tree_limit, approximate, check_additivity, from_call)
479 out = self._get_shap_output(phi, flat_output)
480 if check_additivity and self.model.model_output == "raw":
--> 481 self.assert_additivity(out, self.model.predict(X))
483 return out
File ~/mambaforge-pypy3/envs/arfs/lib/python3.10/site-packages/fasttreeshap/explainers/_tree.py:641, in Tree.assert_additivity(self, phi, model_output)
639 check_sum(self.expected_value[i] + phi[i].sum(-1), model_output[:,i])
640 else:
--> 641 check_sum(self.expected_value + phi.sum(-1), model_output)
File ~/mambaforge-pypy3/envs/arfs/lib/python3.10/site-packages/fasttreeshap/explainers/_tree.py:635, in Tree.assert_additivity.<locals>.check_sum(sum_val, model_output)
631 err_msg += " Consider retrying with the feature_perturbation='interventional' option."
632 err_msg += " This check failed because for one of the samples the sum of the SHAP values" \
633 " was %f, while the model output was %f. If this difference is acceptable" \
634 " you can set check_additivity=False to disable this check." % (sum_val[ind], model_output[ind])
--> 635 raise Exception(err_msg)
Exception: Additivity check failed in TreeExplainer! Please ensure the data matrix you passed to the explainer is the same shape that the model was trained on. If your data shape is correct then please report this on GitHub. Consider retrying with the feature_perturbation='interventional' option. This check failed because for one of the samples the sum of the SHAP values was 239.352906, while the model output was 243.357726. If this difference is acceptable you can set check_additivity=False to disable this check.
FastTreeShap fails when using XGBoost, I opened an issue.
[25]:
import fasttreeshap
import shap
import xgboost
print(
f"Using xgboost {xgboost.__version__}, shap {shap.__version__} and fasttreeshap {fasttreeshap.__version__}"
)
Using xgboost 1.7.6, shap 0.42.1 and fasttreeshap 0.1.6
Testing GrootCV#
[26]:
# Testing the changes with rnd cat. and num. predictors added to the set of genuine predictors
def testing_estimators(X, y, sample_weight=None, objective="rmse"):
feat_selector = arfsgroot.GrootCV(
objective=objective, cutoff=1, n_folds=5, n_iter=5, fastshap=False
)
feat_selector.fit(X, y, sample_weight)
print(feat_selector.get_feature_names_out())
fig = feat_selector.plot_importance(n_feat_per_inch=5)
# highlight synthetic random variable
fig = highlight_tick(figure=fig, str_match="random")
fig = highlight_tick(figure=fig, str_match="genuine", color="green")
plt.show()
gc.enable()
del feat_selector
gc.collect()
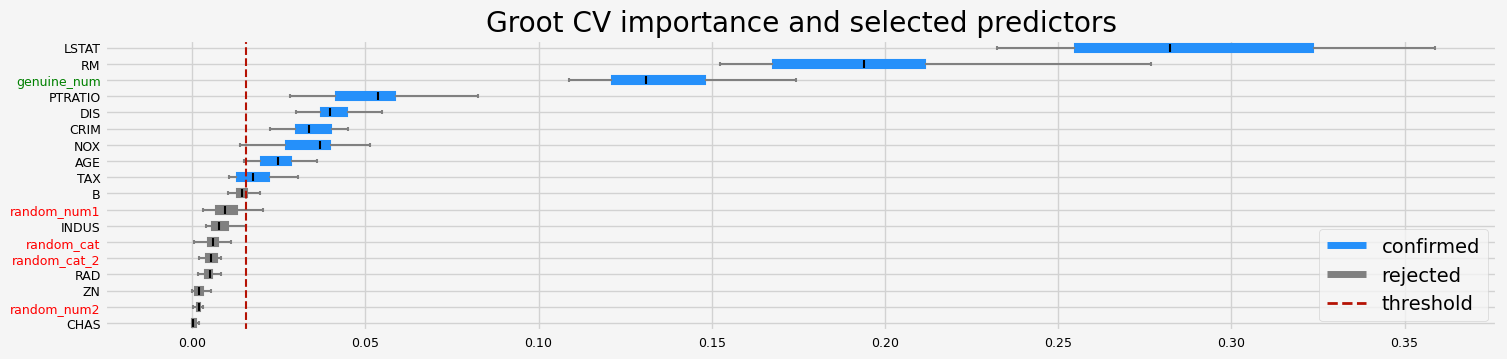
if __name__ == "__main__":
# regression
boston = load_data(name="Boston")
X, y = boston.data, boston.target
cat_f = boston.categorical
testing_estimators(X=X, y=y, objective="rmse")
['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'TAX' 'PTRATIO' 'LSTAT' 'genuine_num']
[27]:
# Testing the changes with rnd cat. and num. predictors added to the set of genuine predictors
def testing_estimators(X, y, sample_weight=None, objective="rmse"):
feat_selector = arfsgroot.GrootCV(
objective=objective, cutoff=1, n_folds=5, n_iter=5, fastshap=True
)
feat_selector.fit(X, y, sample_weight)
print(feat_selector.get_feature_names_out())
fig = feat_selector.plot_importance(n_feat_per_inch=5)
# highlight synthetic random variable
fig = highlight_tick(figure=fig, str_match="random")
fig = highlight_tick(figure=fig, str_match="genuine", color="green")
plt.show()
gc.enable()
del feat_selector
gc.collect()
if __name__ == "__main__":
# regression
boston = load_data(name="Boston")
X, y = boston.data, boston.target
cat_f = boston.categorical
testing_estimators(X=X, y=y, objective="rmse")
['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'TAX' 'PTRATIO' 'LSTAT' 'genuine_num']
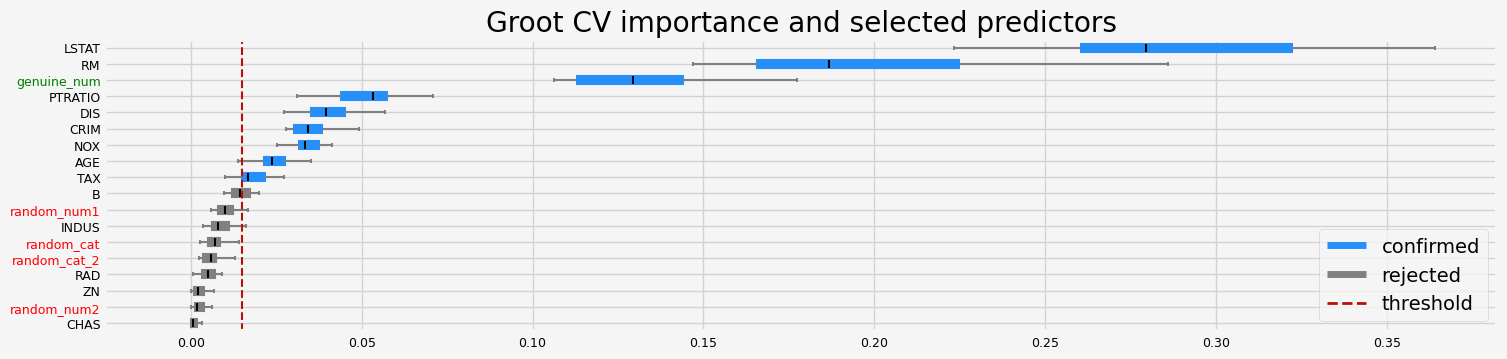
Testing BoostAGroota#
[28]:
models = [
RandomForestRegressor(n_jobs=4, oob_score=True),
CatBoostRegressor(random_state=42, verbose=0),
LGBMRegressor(random_state=42, verbose=-1),
LightForestRegressor(n_feat=X.shape[1]),
XGBRegressor(random_state=42, verbosity=0),
]
feat_selector = arfsgroot.BoostAGroota(
estimator=model, cutoff=1, iters=10, max_rounds=10, delta=0.1
)
if __name__ == "__main__":
# regression
boston = load_data(name="Boston")
X, y = boston.data, boston.target
cat_f = boston.categorical
# running the ARFS methods using different models
compare_varimp(feat_selector, models, X, y, sample_weight=None)
==================== BoostAGroota - testing: RandomForestRegressor for var.imp: shap ====================
['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'TAX' 'PTRATIO' 'B' 'LSTAT' 'random_num1'
'genuine_num']
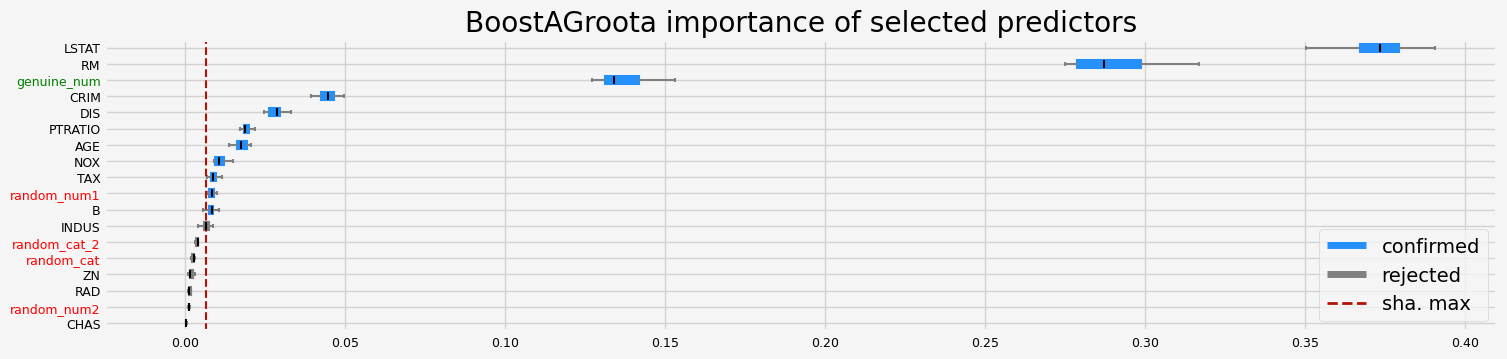
==================== BoostAGroota - testing: RandomForestRegressor for var.imp: fastshap ====================
['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'TAX' 'PTRATIO' 'B' 'LSTAT' 'random_num1'
'genuine_num']
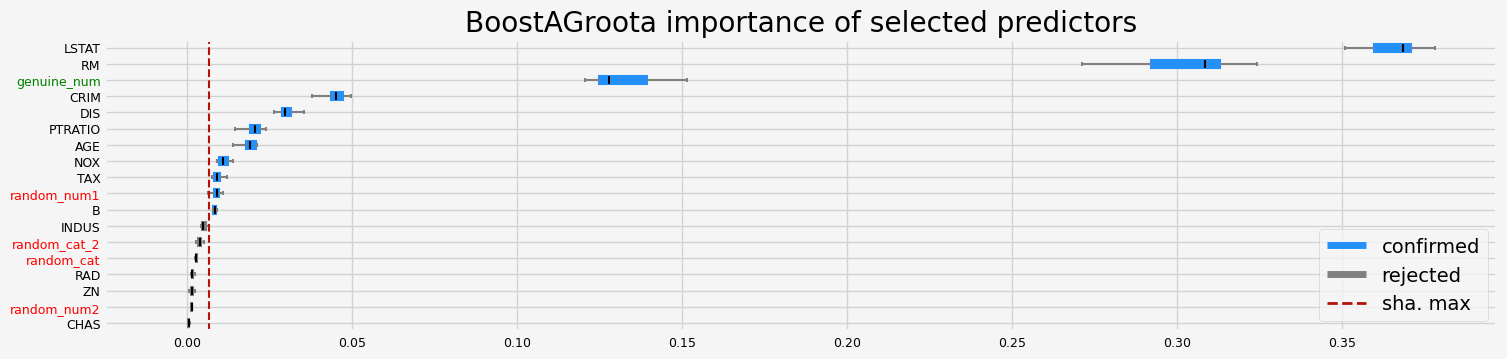
==================== BoostAGroota - testing: RandomForestRegressor for var.imp: pimp ====================
['CRIM' 'INDUS' 'NOX' 'RM' 'AGE' 'DIS' 'TAX' 'PTRATIO' 'LSTAT'
'genuine_num']

==================== BoostAGroota - testing: RandomForestRegressor for var.imp: native ====================
['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'TAX' 'PTRATIO' 'B' 'LSTAT' 'genuine_num']
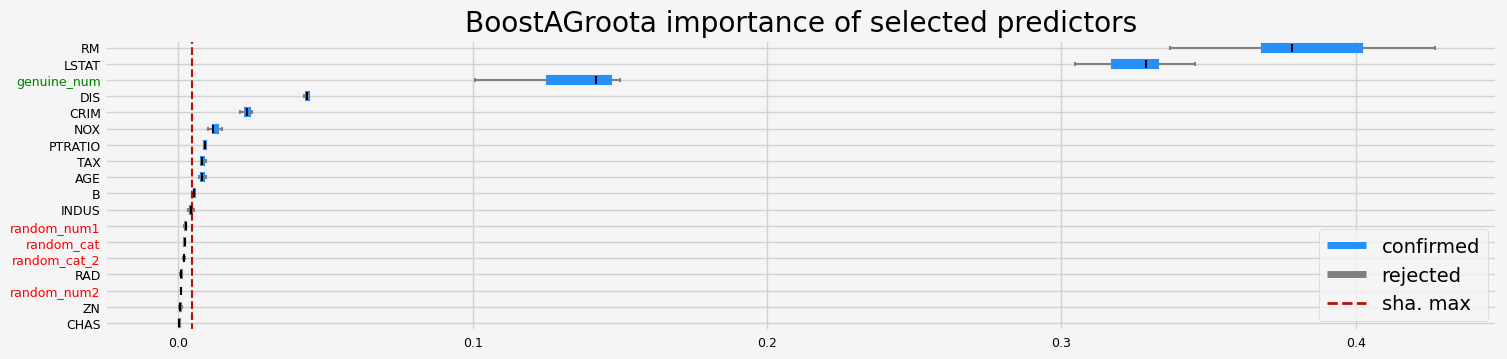
==================== BoostAGroota - testing: CatBoostRegressor for var.imp: shap ====================
['CRIM' 'INDUS' 'NOX' 'RM' 'AGE' 'DIS' 'TAX' 'PTRATIO' 'B' 'LSTAT'
'genuine_num']

==================== BoostAGroota - testing: CatBoostRegressor for var.imp: fastshap ====================
['CRIM' 'ZN' 'INDUS' 'NOX' 'RM' 'AGE' 'DIS' 'TAX' 'PTRATIO' 'B' 'LSTAT'
'genuine_num']

==================== BoostAGroota - testing: CatBoostRegressor for var.imp: pimp ====================
['CRIM' 'INDUS' 'NOX' 'RM' 'AGE' 'DIS' 'TAX' 'PTRATIO' 'B' 'LSTAT'
'genuine_num']
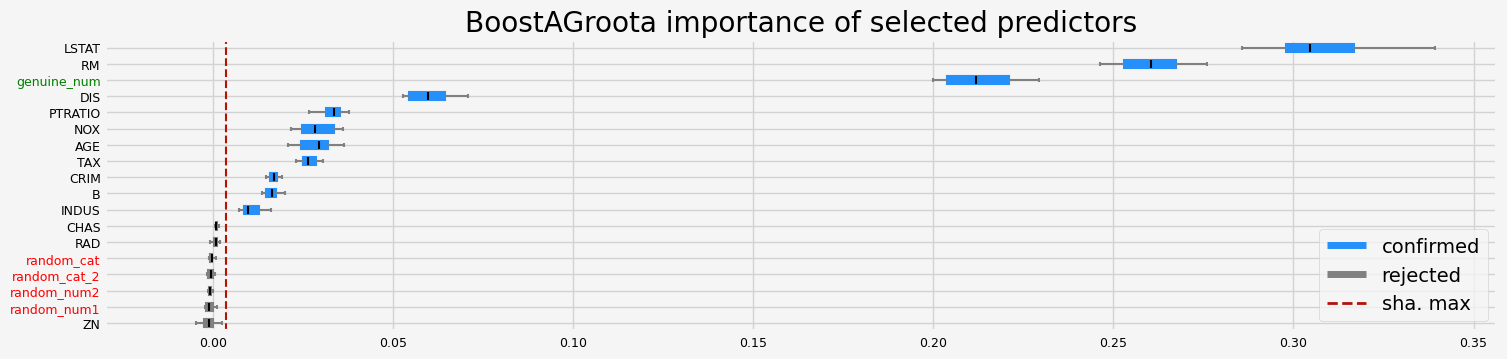
==================== BoostAGroota - testing: CatBoostRegressor for var.imp: native ====================
['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'TAX' 'PTRATIO' 'B' 'LSTAT' 'genuine_num']

==================== BoostAGroota - testing: LGBMRegressor for var.imp: shap ====================
['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'PTRATIO' 'LSTAT' 'genuine_num']
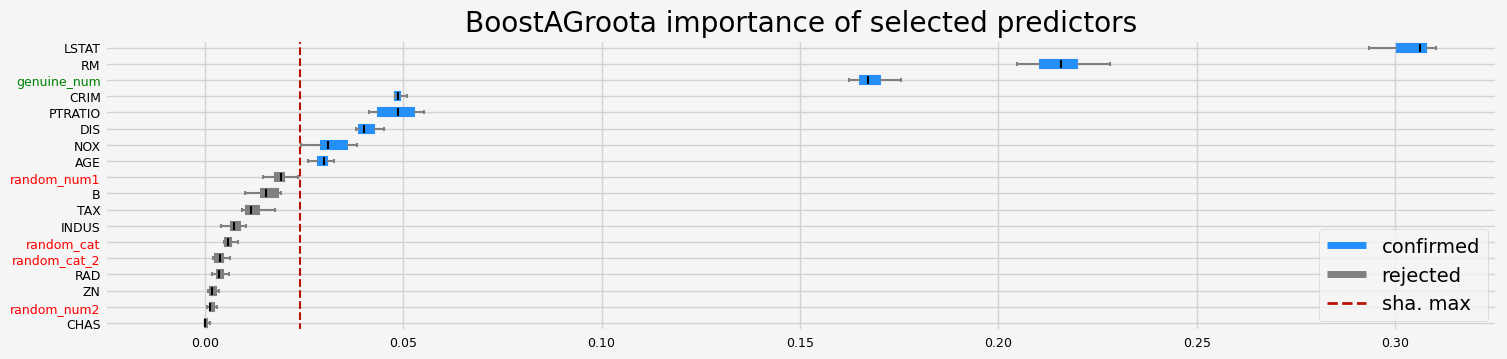
==================== BoostAGroota - testing: LGBMRegressor for var.imp: fastshap ====================
['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'PTRATIO' 'LSTAT' 'genuine_num']
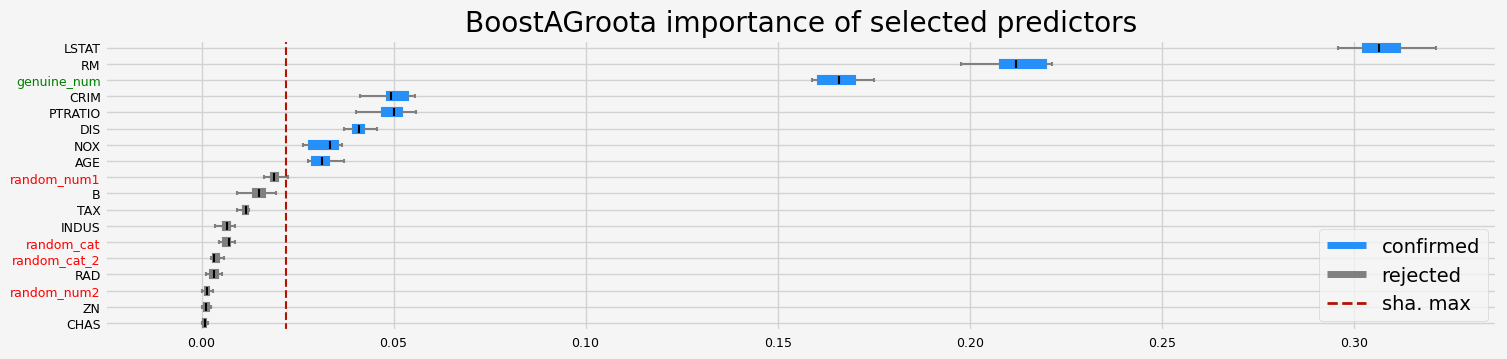
==================== BoostAGroota - testing: LGBMRegressor for var.imp: pimp ====================
['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'TAX' 'PTRATIO' 'LSTAT' 'genuine_num']
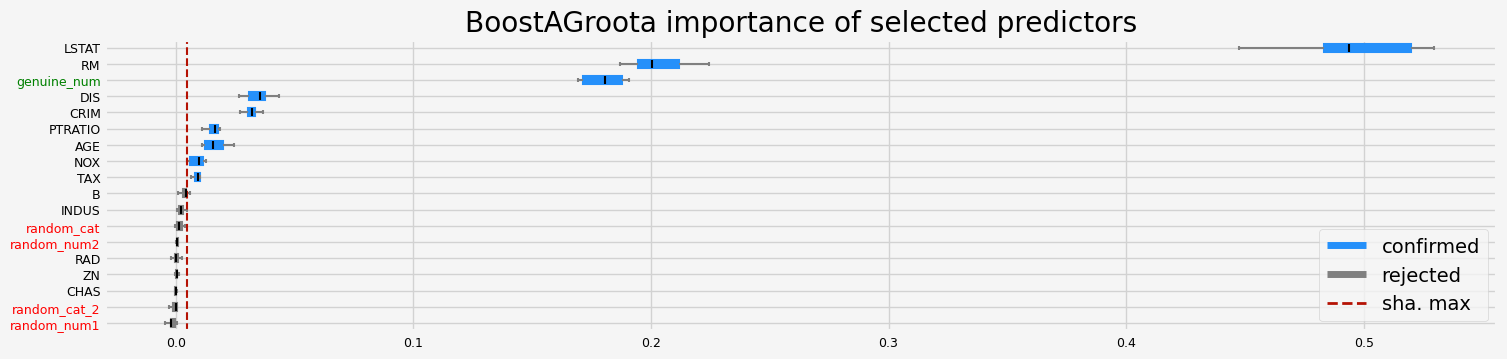
==================== BoostAGroota - testing: LGBMRegressor for var.imp: native ====================
['CRIM' 'RM' 'DIS' 'LSTAT' 'genuine_num']
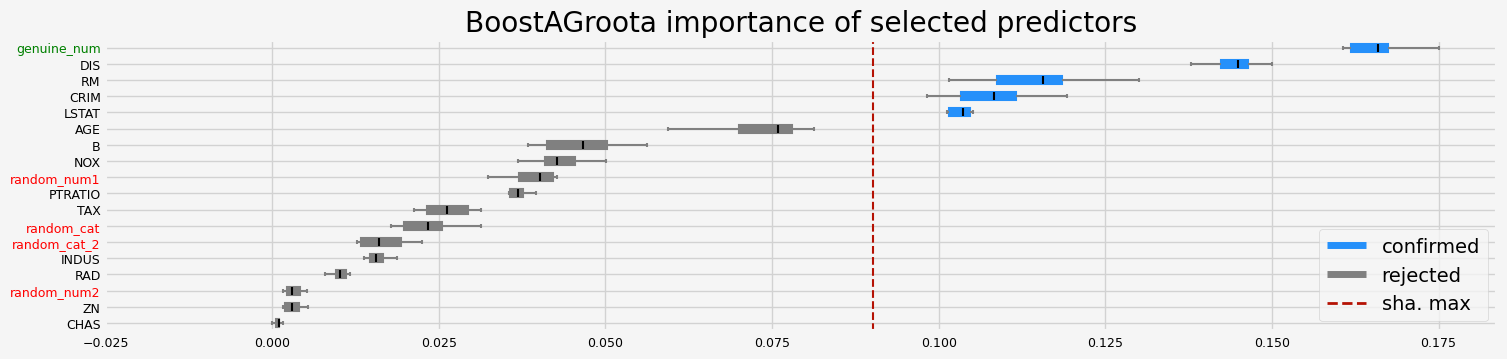
==================== BoostAGroota - testing: LGBMRegressor for var.imp: shap ====================
['CRIM' 'INDUS' 'RM' 'DIS' 'TAX' 'LSTAT' 'genuine_num']
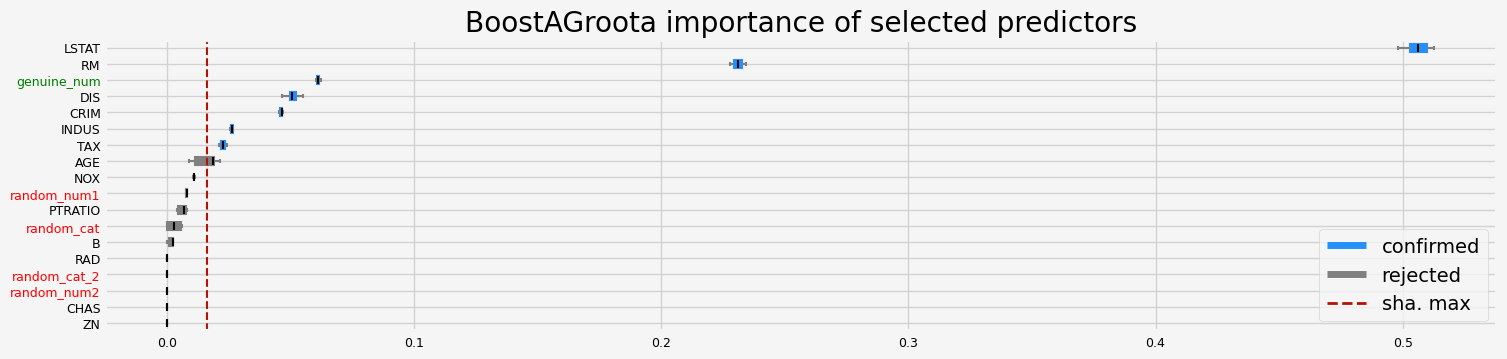
==================== BoostAGroota - testing: LGBMRegressor for var.imp: fastshap ====================
['CRIM' 'INDUS' 'NOX' 'RM' 'AGE' 'DIS' 'TAX' 'LSTAT' 'genuine_num']
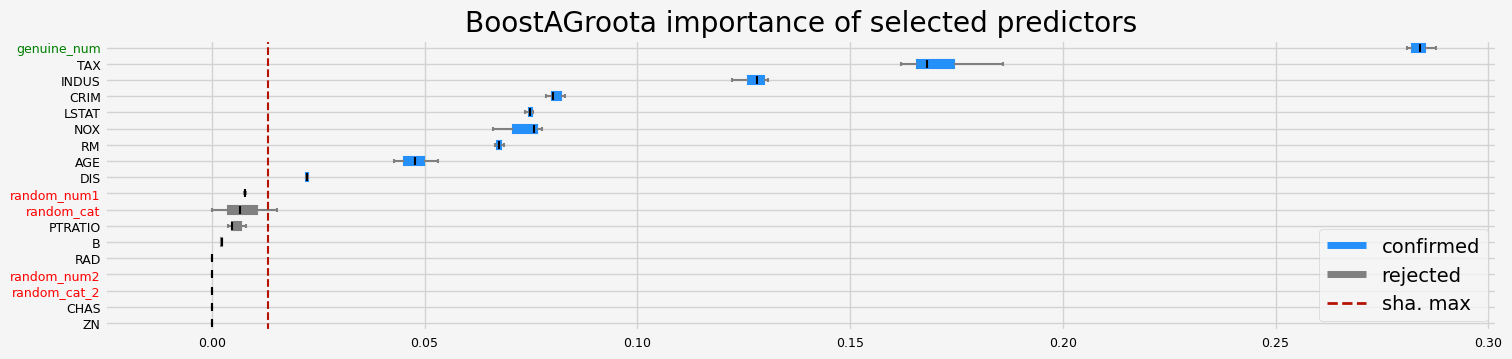
==================== BoostAGroota - testing: LGBMRegressor for var.imp: pimp ====================
['CRIM' 'INDUS' 'NOX' 'RM' 'DIS' 'LSTAT' 'genuine_num']
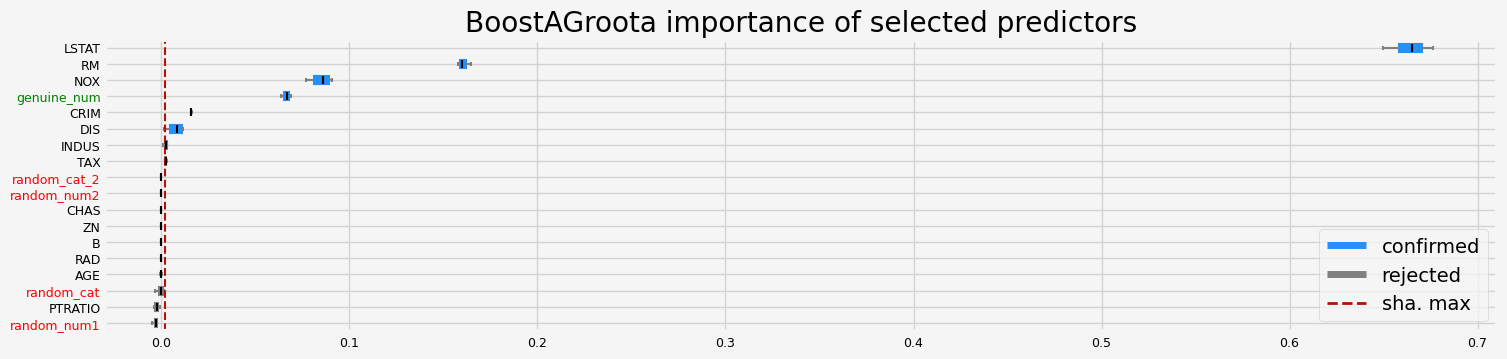
==================== BoostAGroota - testing: LGBMRegressor for var.imp: native ====================
['CRIM']
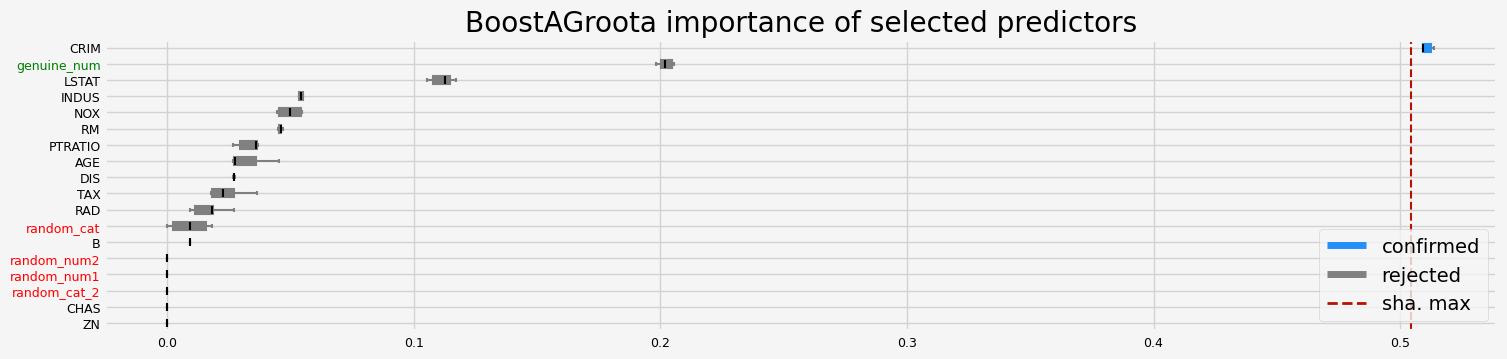
==================== BoostAGroota - testing: XGBRegressor for var.imp: shap ====================
['CRIM' 'NOX' 'RM' 'AGE' 'DIS' 'TAX' 'PTRATIO' 'B' 'LSTAT' 'random_num1'
'genuine_num']
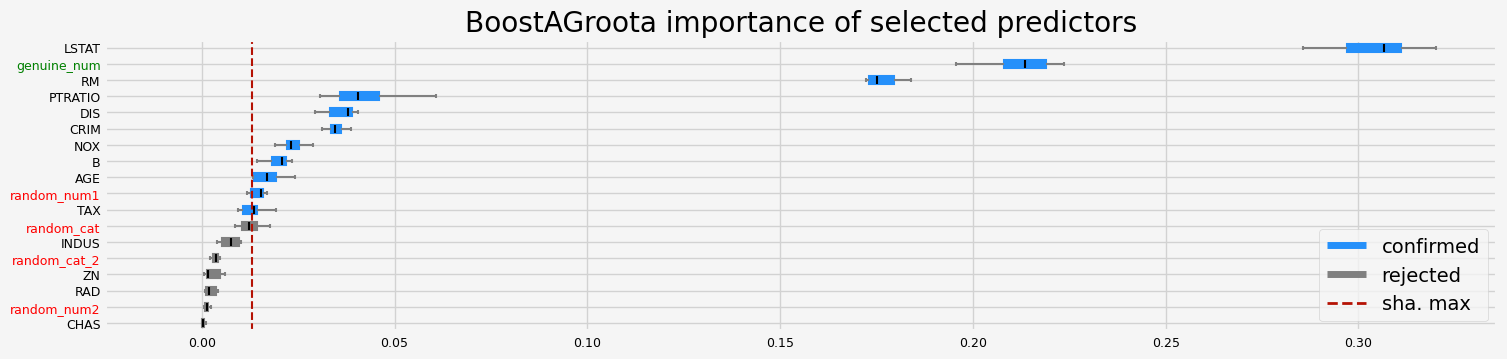
==================== BoostAGroota - testing: XGBRegressor for var.imp: fastshap ====================
---------------------------------------------------------------------------
Exception Traceback (most recent call last)
Cell In[28], line 19
17 cat_f = boston.categorical
18 # running the ARFS methods using different models
---> 19 compare_varimp(feat_selector, models, X, y, sample_weight=None)
File ~/Documents/arfs/src/arfs/benchmark.py:142, in compare_varimp(feat_selector, models, X, y, sample_weight)
140 feat_selector.estimator = mod_clone
141 # fit the feature selector
--> 142 feat_selector.fit(X=X, y=y, sample_weight=sample_weight)
143 # print the results
144 print(feat_selector.selected_features_)
File ~/Documents/arfs/src/arfs/feature_selection/allrelevant.py:1574, in BoostAGroota.fit(self, X, y, sample_weight)
1571 sample_weight = pd.Series(_check_sample_weight(sample_weight, X))
1573 # crit, keep_vars, df_vimp, mean_shadow
-> 1574 _, self.selected_features_, self.sha_cutoff_df, self.mean_shadow = _boostaroota(
1575 X,
1576 y,
1577 # metric=self.metric,
1578 estimator=self.estimator,
1579 cutoff=self.cutoff,
1580 iters=self.iters,
1581 max_rounds=self.max_rounds,
1582 delta=self.delta,
1583 silent=self.silent,
1584 weight=sample_weight,
1585 imp=self.importance,
1586 )
1587 self.selected_features_ = self.selected_features_.values
1588 self.support_ = np.asarray(
1589 [c in self.selected_features_ for c in self.feature_names_in_]
1590 )
File ~/Documents/arfs/src/arfs/feature_selection/allrelevant.py:1877, in _boostaroota(X, y, estimator, cutoff, iters, max_rounds, delta, silent, weight, imp)
1874 while True:
1875 # Inside this loop we reduce the dataset on each iteration exiting with keep_vars
1876 i += 1
-> 1877 crit, keep_vars, df_vimp, mean_shadow = _reduce_vars_sklearn(
1878 new_x,
1879 y,
1880 estimator=estimator,
1881 this_round=i,
1882 cutoff=cutoff,
1883 n_iterations=iters,
1884 delta=delta,
1885 silent=silent,
1886 weight=weight,
1887 imp_kind=imp,
1888 cat_feature=cat_idx,
1889 )
1891 b_df = df_vimp.T.iloc[1:-1].astype(float)
1892 b_df.columns = df_vimp.T.iloc[0].values
File ~/Documents/arfs/src/arfs/feature_selection/allrelevant.py:1774, in _reduce_vars_sklearn(X, y, estimator, this_round, cutoff, n_iterations, delta, silent, weight, imp_kind, cat_feature)
1767 new_x, shadow_names = _create_shadow(X)
1768 imp_func = {
1769 "shap": _get_shap_imp,
1770 "fastshap": _get_shap_imp_fast,
1771 "pimp": _get_perm_imp,
1772 "native": _get_imp,
1773 }
-> 1774 importance = imp_func[imp_kind](
1775 estimator, new_x, y, sample_weight=weight, cat_feature=cat_feature
1776 )
1779 # Create a dataframe to store the feature importances
1780 if i == 1:
File ~/Documents/arfs/src/arfs/feature_selection/allrelevant.py:1302, in _get_shap_imp_fast(estimator, X, y, sample_weight, cat_feature)
1293 model, X_tt, y_tt, w_tt = _split_fit_estimator(
1294 estimator, X, y, sample_weight=sample_weight, cat_feature=cat_feature
1295 )
1296 explainer = FastTreeExplainer(
1297 model,
1298 algorithm="auto",
1299 shortcut=False,
1300 feature_perturbation="tree_path_dependent",
1301 )
-> 1302 shap_matrix = explainer.shap_values(X_tt)
1303 # multiclass returns a list
1304 # for binary and for some models, shap is still returning a list
1305 if is_classifier(estimator):
File ~/mambaforge-pypy3/envs/arfs/lib/python3.10/site-packages/fasttreeshap/explainers/_tree.py:481, in Tree.shap_values(self, X, y, tree_limit, approximate, check_additivity, from_call)
479 out = self._get_shap_output(phi, flat_output)
480 if check_additivity and self.model.model_output == "raw":
--> 481 self.assert_additivity(out, self.model.predict(X))
483 return out
File ~/mambaforge-pypy3/envs/arfs/lib/python3.10/site-packages/fasttreeshap/explainers/_tree.py:641, in Tree.assert_additivity(self, phi, model_output)
639 check_sum(self.expected_value[i] + phi[i].sum(-1), model_output[:,i])
640 else:
--> 641 check_sum(self.expected_value + phi.sum(-1), model_output)
File ~/mambaforge-pypy3/envs/arfs/lib/python3.10/site-packages/fasttreeshap/explainers/_tree.py:635, in Tree.assert_additivity.<locals>.check_sum(sum_val, model_output)
631 err_msg += " Consider retrying with the feature_perturbation='interventional' option."
632 err_msg += " This check failed because for one of the samples the sum of the SHAP values" \
633 " was %f, while the model output was %f. If this difference is acceptable" \
634 " you can set check_additivity=False to disable this check." % (sum_val[ind], model_output[ind])
--> 635 raise Exception(err_msg)
Exception: Additivity check failed in TreeExplainer! Please ensure the data matrix you passed to the explainer is the same shape that the model was trained on. If your data shape is correct then please report this on GitHub. Consider retrying with the feature_perturbation='interventional' option. This check failed because for one of the samples the sum of the SHAP values was -13.068551, while the model output was 9.339367. If this difference is acceptable you can set check_additivity=False to disable this check.
[ ]: